Cloud-Based Phrase Mining and Analysis of User-Defined Phrase-Category Association in Biomedical Publications
Özet
We present a protocol and associated programming code as well as metadata samples to support a cloud-based automated identification of phrases-category association representing unique concepts in user selected knowledge domain in biomedical literature. The phrase-category association quantified by this protocol can facilitate in depth analysis in the selected knowledge domain.
Abstract
The rapid accumulation of biomedical textual data has far exceeded the human capacity of manual curation and analysis, necessitating novel text-mining tools to extract biological insights from large volumes of scientific reports. The Context-aware Semantic Online Analytical Processing (CaseOLAP) pipeline, developed in 2016, successfully quantifies user-defined phrase-category relationships through the analysis of textual data. CaseOLAP has many biomedical applications.
We have developed a protocol for a cloud-based environment supporting the end-to-end phrase-mining and analyses platform. Our protocol includes data preprocessing (e.g., downloading, extraction, and parsing text documents), indexing and searching with Elasticsearch, creating a functional document structure called Text-Cube, and quantifying phrase-category relationships using the core CaseOLAP algorithm.
Our data preprocessing generates key-value mappings for all documents involved. The preprocessed data is indexed to carry out a search of documents including entities, which further facilitates the Text-Cube creation and CaseOLAP score calculation. The obtained raw CaseOLAP scores are interpreted using a series of integrative analyses, including dimensionality reduction, clustering, temporal, and geographical analyses. Additionally, the CaseOLAP scores are used to create a graphical database, which enables semantic mapping of the documents.
CaseOLAP defines phrase-category relationships in an accurate (identifies relationships), consistent (highly reproducible), and efficient manner (processes 100,000 words/sec). Following this protocol, users can access a cloud-computing environment to support their own configurations and applications of CaseOLAP. This platform offers enhanced accessibility and empowers the biomedical community with phrase-mining tools for widespread biomedical research applications.
Introduction
Manual evaluation of millions of text files for the study of phrase-category association (e.g., Age group to protein association) is incomparable with the efficiency provided by an automated computational method. We want to introduce the cloud-based Context-aware Semantic Online Analytical Processing (CaseOLAP) platform as a phrase-mining method for automated computation of phrase-category association in the biomedical context.
The CaseOLAP platform, which was first defined in 20161, is very efficient compared to the traditional methods of data management and computation because of its functional document management called Text-Cube2,3,4, which distributes the documents while maintaining underlying hierarchy and neighbourhoods. It has been applied in biomedical research5 to study entity-category association. The CaseOLAP platform consists of six major steps including download and extraction of data, parsing, indexing, Text-Cube creation, entity count, and CaseOLAP score calculation; which is the main focus of the protocol (Figure 1, Figure 2, Table 1).
To implement the CaseOLAP algorithm, the user sets up categories of interest (e.g., disease, signs and symptoms, age groups, diagnosis) and entities of interest (e.g., proteins, drugs). One example of a category included in this article is the 'Age Groups', which has 'infant', 'child', 'adolescent', and 'adult' subcategories as cells of the Text-Cube and protein names (synonyms) and abbreviations as entities. Medical Subject Headings (MeSH) are implemented to retrieve publications corresponding to the defined categories (Table 2). MeSH descriptors are organized in a hierarchical tree structure to permit search for publications at varying levels of specificity (a sample shown in Figure 3). The CaseOLAP platform utilizes the data indexing and search functionality for curation of the documents associated with an entity which further facilitate document to entity count mapping and CaseOLAP score calculation.
The details of the CaseOLAP score calculation is available in previous publications1,5. This score is computed using specific ranking criteria based on underlying Text-Cube document structure. The final score is the product of Integrity, Popularity, and Distinctiveness. Integrity describes whether a representative entity is an integral semantic unit that collectively refers to a meaningful concept. The integrity of the user-defined phrase is taken to be 1.0 because it stands as a standard phrase in the literature. Distinctiveness represents the relative relevance of a phrase in one subset of documents compared to the rest of the other cells. It first calculates the relevance of an entity to a specific cell by comparing the occurrence of the protein name in the target data set and provides a normalized Distinctiveness score. Popularity represents the fact that phrase with a higher popularity score appears more frequently in one subset of documents. Rare protein names in a cell are ranked low, while an increase in their frequency of mention has a diminishing return due to the implementation of the logarithmic function of frequency. Quantitatively measuring these three concepts depends on the (1) term frequency of the entity over a cell and across the cells and (2) number of documents having that entity (document frequency) within the cell and across the cells.
We have studied two representative scenarios using a PubMed dataset and our algorithm. We are interested in how mitochondrial proteins are associated with two unique categories of MeSH descriptors; "Age Groups" and "Nutritional and Metabolic Diseases". Specifically, we retrieved 15,728,250 publications from 20 years publications collected by PubMed (1998 to 2018), among them, 8,123,458 unique abstracts have had full MeSH descriptors. Accordingly, 1,842 human mitochondrial protein names (including abbreviations and synonyms), acquired from UniProt (uniprot.org) as well as from MitoCarta2.0 (http://mitominer.mrc-mbu.cam.ac.uk/release-4.0/begin.do>), are systematically examined. Their associations with these 8,899,019 publications and entities were studied using our protocol; we constructed a Text-Cube and calculated the respective CaseOLAP scores.
Protocol
NOTE: We have developed this protocol based on the Python programming language. To run this program, have Anaconda Python and Git pre-installed on the device. The commands provided in this protocol are based on Unix environment. This protocol provides the detail of downloading data from PubMed (MEDLINE) database, parsing the data, and setting up a cloud computing platform for the phrase mining and quantification of user-defined entity-category association.
1.Getting code and python environment setup
- Download or clone the code repository from Github (https://github.com/CaseOLAP/caseolap) or by typing ‘git clone https://github.com/CaseOLAP/caseolap.git’ in the terminal window.
- Navigate to the ‘caseolap’ directory. This is the root directory of the project. Within this directory, the ‘data’ directory will be populated with multiple data sets as you progress through these steps in the protocol. The ‘input’ directory is for user-provided data. The ‘log’ directory has log files for troubleshooting purposes. The ‘result’ directory is where the final results will be stored.
- Using the terminal window, go to the directory where you cloned our GitHub repository. Create the CaseOLAP environment using the ‘environment.yml’ file by typing ‘conda env create -f environment.yaml’ in the terminal. Then activate the environment by typing ‘source activate caseolap’ in the terminal.
2. Downloading documents
- Make sure that the FTP address in ‘ftp_configuration.json’ in the config directory is the same as the Annual Baseline or Daily Update Files link address, found in the link (https://www.nlm.nih.gov/databases/download/pubmed_medline.html).
- To download baseline only or update files only, set ‘true’ in the ‘download_config.json’ file in the ‘config’ directory. By default, it downloads and extracts both baseline and update files. A sample of extracted XML data can be viewed at (https://github.com/CaseOLAP/caseolap-pipelines/blob/master/data/extracted-data-sample.xml)
- Type ‘python run_download.py’ in the terminal window to download abstracts from the Pubmed database. This will create a directory called ‘ftp.ncbi.nlm.nih.gov’ in the current directory. This process checks the integrity of the downloaded data and extracts it to the target directory.
- Go to the ‘log’ directory to read the log messages in ‘download_log.txt’ in case the download process fails. If the process is completed successfully, the debugging messages of the download process will be printed out in this log file.
- When the download is complete, navigate through ‘ftp.ncbi.nlm.nih.gov’ to make sure that there is ‘updatefiles’ or ‘basefiles’ or both directories based on download configuration in ‘download_config.json’. The file statistics become available at ‘filestat.txt’ in the ‘data’ directory.
3. Parsing documents
- Make sure that downloaded and extracted data is available at ‘ftp.ncbi.nlm.nih.gov’ directory from step 2. This directory is the input data directory in this step.
- To modify the data-parsing schema, select parameters in ‘parsing_config.json’ file in the ‘config’ directory by setting their value to ‘true’. By default, it parses the PMID, authors, abstract, MeSH, location, journal, publication date.
- Type ‘python run_parsing.py’ in the terminal to parse the documents from downloaded (or extracted) files. This step parses all downloaded XML files and creates a python dictionary for each document with keys (e.g., PMID, authors, abstract, MeSH of the file based on parsing schema setup at step 3.2).
- Once data parsing is completed, make sure that parsed data is saved in the file called ‘pubmed.json’ in the data directory. A sample of parsed data is available at Figure 3.
- Go to the ‘log’ directory to read the log messages in ‘parsing_log.txt’ in case the parsing process fails. If the process is completed successfully, the debugging messages will be printed out in the log file.
4. Mesh to PMID mapping
- Make sure that parsed data (‘pubmed.json’) is available at the ‘data’ directory.
- Type ‘python run_mesh2pmid.py’ in the terminal to perform MeSH to PMID mapping. This creates a mapping table where each of the MeSH collects associated PMIDs. A Single PMID may fall under the multiple MeSH terms.
- Once the mapping is completed, make sure that there is ‘mesh2pmid.json’ in the data directory. A sample of the top 20 mapping statistics is available in Table-2, Figures 4 and 5.
- Go to the ‘log’ directory to read the log messages in ‘mesh2pmid_mapping_log.txt’ in case this process fails. If the process is completed successfully, the debugging messages of the mapping will be printed out in this log file.
5. Document indexing
- Download the Elasticsearch application from https://www.elastic.co. Currently, the download is available at (https://www.elastic.co/downloads/elasticsearch). To download the software in the remote cloud, type ‘wget https://artifacts.elastic.co/downloads/elasticsearch/elasticsearch-x.x.x.tar.gz’ in the terminal. Make sure that ‘x.x.x’ in the above command is replaced by proper version number.
- Make sure that downloaded ‘elasticsearch-x.x.x.tar.gz’ file appears in the root directory then extract the files by typing ‘tar xvzf elasticsearch-x.x.x.tar.gz’ in the terminal window.
- Open a new terminal and go to the ElasticSearch bin directory by typing ‘cd Elasticsearch/bin’ in the terminal from the root directory.
- Start the Elasticsearch server by typing ‘./Elasticsearch’ in the terminal window. Make sure that the server is started without error messages. In case of error on starting Elasticsearch server, follow the instructions at (https://www.elastic.co/guide/en/elasticsearch/reference/current/index.html).
- Modify the contents in the ‘index_init_config.json’ in ‘config’ directory to set index initiation. By default, it will select all items present.
- Type ‘python run_index_init.py’ in the terminal to initiate an index-database in the Elasticsearch server. This initializes the index with a set of criteria known as index information (e.g., index name, type name, number of shards, number of replicas). You will see the message mentioning index is successfully created.
- Select the items in the ‘index_populate_config.json’ in the ‘config’ directory by setting their value to ‘true’. By default, it will select all items present.
- Make sure that parsed data (‘pubmed.json’) is present in the ‘data’ directory.
- Type ‘python run_index_populate.py’ in the terminal to populate the index by creating bulk data with two components.A first component is a dictionary with metadata information on the index name, type name, and bulk id (e.g., ‘PMID’). A second component is a data dictionary containing all the information on the tags (e.g., ‘title’,’abstract’,’MeSH’).
- Go to the ‘log’ directory to read the log messages in ‘indexing_log.txt’ in case this process fails. If the process is completed successfully, the debugging messages of the indexing will be printed out in the log file.
6. Text-cube creation
- Download the latest MeSH Tree available at (https://www.nlm.nih.gov/mesh/filelist.html). The current version of the code is using MeSH Tree 2018 as ‘meshtree2018.bin’ in the input directory.
- Define the categories of interest (e.g., Disease names, Age groups, Gender). A category may include one or more MeSH descriptors (https://meshb-prev.nlm.nih.gov/treeView). Collect MeSH IDs for a category. Save the names of the categories in the file ‘textcube_config.json’ in the config directory (see a sample of the category in ‘Age Group’ in the downloaded version of ‘textcube_config.json’ file).
- Put the collected categories of MeSH IDs in a line separated by a space. Save the category file as ‘categories.txt’ in the ‘input’ directory (see a sample of ‘Age Group’ MeSH IDs in the downloaded version of ‘categories.txt’ file). This algorithm automatically selects all descendent MeSH descriptors. An example of root nodes and descendants are presented in Figure 4.
- Make sure that ‘mesh2pmid.json’ is in the ‘data’ directory. If the MeSH Tree has been updated with a different name (e.g., ‘meashtree2019.bin’) in ‘input’ directory, make sure that this is properly represented in the input data path in the ‘run_textube.py’ file.
- Type ‘python run_textcube.py’ in the terminal to create a document data structure called Text-Cube. This creates a collection of documents (PMIDs) for each category. A single document (PMID) may fall under multiple categories, (see Table 3A, Table 3B, Figure 6A and Figure 7A).
- Once Text-Cube creation step is completed, make sure that following data files are saved in the ‘data’ directory: (1) a cell to PMID table as “textcube_cell2pmid.json”, (2) a PMID to cell mapping table as “textcube_pmid2cell.json”, (3) a collection of all descendant MeSH terms for a cell as “meshterms_per_cat.json” (4) Text-Cube data statistics as “textcube_stat.txt”.
- Go to the ‘log’ directory to read the log messages in ‘textcube_log.txt’ in case this process fails. If the process is completed successfully, the debugging messages of the Text-Cube creation will be printed out in the log file.
7. Entity count
- Create user-defined entities (e.g., protein names, genes, chemicals). Put one entity and its abbreviations in a single line separated by “|”. Save the entity file as ‘entities.txt’ in the ‘input’ directory. A sample of entities can be found in Table 4.
- Make sure that Elasticsearch server is running. Otherwise, go to step 5.2 and 5.3 to restart the Elasticsearch server. It is expected to have an indexed database called ‘pubmed’ in your Elasticsearch server which was established in step 5.
- Make sure that ‘textcube_pmid2cell.json’ is in the ‘data’ directory.
- Type ‘python run_entitycount.py’ in the terminal to perform Entity Count operation. This searches the documents from the indexed database and counts the entity in each document as well as collects the PMIDs in which entities were found.
- Once the Entity Count is completed, make sure that the final results are saved as ‘entitycount.txt’ and ‘entityfound_pmid2cell.json’ in the ‘data’ directory.
- Go to the ‘log’ directory to read the log messages in ‘entitycount_log.txt’ in case this process fails. If the process is completed successfully, the debugging messages of the Entity Count will be printed out in the log file.
8. Metadata update
- Make sure that all input data (‘entitycount.txt’, ‘textcube_pmid2cell.json’, ‘entityfound_pmid2cell.txt’) are in the ‘data’ directory. These are the input data for Metadata Update.
- Type ‘python run_metadata_update.py’ in the terminal to update the metadata. This prepares a collection of metadata (e.g., cell name, associated MeSH, PMIDs) representing each text document in the cell. A sample of Text-Cube metadata is presented in Table 3A and Table 3B.
- Once the Metadata Update is completed, make sure that ‘metadata_pmid2pcount.json’ and ‘metadata_cell2pmid.json’ files are saved in ‘data’ directory.
- Go to the ‘log’ directory to read the log messages in ‘metadata_update_log.txt’ in case this process fails. If the process is completed successfully, the debugging messages of the metadata update will be printed out in the log file.
9. CaseOLAP score calculation
- Make sure that ‘metadata_pmid2pcount.json’ and ‘metadata_cell2pmid.json’ files are present in the ‘data’ directory. These are the input data for score calculation.
- Type ‘python run_caseolap_score.py’ in the terminal to perform CaseOLAP score calculation. This calculates the CaseOLAP score of the entities based on user-defined categories. The CaseOLAP score is the product of Integrity, Popularity, and Distinctiveness.
- Once the score computation is completed, make sure that this saves the results in multiple files (e.g., popularity as ‘pop.csv’, distinctiveness as ‘dist.csv’, CaseOLAP score as ’caseolap.csv’), in the ‘result’ directory. The summary of the CaseOLAP score calculation is also presented in Table 5.
- Go to the ‘log’ directory to read the log messages in ‘caseolap_score_log.txt’ in case this process fails. If the process is completed successfully, the debugging messages of the CaseOLAP score calculation will be printed out in the log file.
Representative Results
To produce sample results, we implemented the CaseOLAP algorithm in two subject headings/descriptors: "Age Groups" and "Nutritional and Metabolic Diseases" as use cases.
Age Groups. We selected all 4 subcategories of "Age Groups" (infant, child, adolescent, and adult) as cells in a Text-Cube. The obtained metadata and statistics are shown in Table 3A. The comparison of the number of documents among the Text-Cube cells is displayed in Figure 6A. Adult contains 172,394 documents which is the highest number across all cells. The adult and adolescent subcategories have the highest number of shared documents (26,858 documents). Notably, these documents included the entity of our interest only (i.e., mitochondrial proteins). The Venn diagram in Figure 6B represents the number of entities (i.e., mitochondrial proteins) found within each cell, and within multiple overlaps among the cells. The number of proteins shared within all Age Groups subcategories is 162. The adult subcategory depicts the highest number of unique proteins (151) followed by child (16), infant (8) and adolescent (1). We calculated the protein-age group association as a CaseOLAP score. The top 10 proteins (based on their average CaseOLAP score) associated with infant, child, adolescent and adult subcategories are Sterol 26-hydroxylase, Alpha-crystallin B chain, 25-hydroxyvitamin D-1 alpha-hydroxylase, Serotransferrin, Citrate synthase, L-seryl-tRNA, Sodium/potassium-transporting ATPase subunit alpha-3, Glutathione S-transferase omega-1, NADPH:adrenodoxin oxidoreductase, and Mitochondrial peptide methionine sulfoxide reductase (shown in Figure 6C). The adult subcategory displays 10 heatmap cells with a higher intensity compared to the heatmap cells of the adolescent, child and infant subcategory, indicating that the top 10 mitochondrial proteins exhibit the strongest associations to the adult subcategory. The mitochondrial protein Sterol 26-hydroxylase has high associations in all age subcategories which is demonstrated by heatmap cells with higher intensities compared to the heatmap cells of the other 9 mitochondrial proteins. The statistical distribution of the absolute difference in the score between two groups shows the following range for mean difference with a 99% confidence interval: (1) the mean difference between 'ADLT' and 'INFT' lies in the range (0.029 to 0.042), (2) the mean difference between 'ADLT' and 'CHLD' lies in the range (0.021 to 0.030), (3) the mean difference between 'ADLT' and 'ADOL' lies in the range (0.020 to 0.029), (4) the mean difference between 'ADOL' and 'INFT' lies in the range (0.015 to 0.022), (5) the mean difference between 'ADOL' and 'CHLD' lies in the range (0.007 to 0.010), (6) the mean difference between 'CHLD' and 'INFT' lies in the range (0.011 to 0.016).
Nutritional and Metabolic Diseases. We selected 2 subcategories of "Nutritional and Metabolic Diseases" (i.e., metabolic disease and nutritional disorders) to create 2 cells in a Text-Cube. The obtained metadata and statistics are shown in Table 3B. The comparison of the number of documents among the Text-Cube cells is displayed in Figure 7A. The subcategory metabolic disease contains 54,762 documents followed by 19,181 documents in nutritional disorders. The subcategories metabolic disease and nutritional disorders have 7,101 shared documents. Notably, these documents included the entity of our interest only (i.e., mitochondrial proteins). The Venn diagram in Figure 7B represents the number of entities found within each cell, and within multiple overlaps between the cells. We calculated the protein-"Nutritional and Metabolic Diseases" association as a CaseOLAP score. The top 10 proteins (based on their average CaseOLAP score) associated with this use case are Sterol 26-hydroxylase, Alpha-crystallin B chain, L-seryl-tRNA, Citrate synthase, tRNA pseudouridine synthase A, 25-hydroxyvitamin D-1 alpha-hydroxylase, Glutathione S-transferase omega-1, NADPH: adrenodoxin oxidoreductase, Mitochondrial peptide methionine sulfoxide reductase, Plasminogen activator inhibitor 1 (shown in Figure 7C). More than half (54%) of all proteins are shared between the subcategories metabolic diseases and nutritional disorders (397 proteins). Interestingly, almost half (43%) of all associated proteins in the metabolic disease subcategory are unique (300 proteins), whereas nutritional disorders exhibit only a few unique proteins (35). Alpha-crystallin B chain displays the strongest association to the subcategory metabolic diseases. Sterol 26-hydroxylase, mitochondrial displays the strongest association in the nutritional disorders subcategory, indicating that this mitochondrial protein is highly relevant in studies describing nutritional disorders. The statistical distribution of the absolute difference in the score between two groups 'MBD' and 'NTD' shows the range (0.046 to 0.061) for the mean difference as a 99% confidence interval.
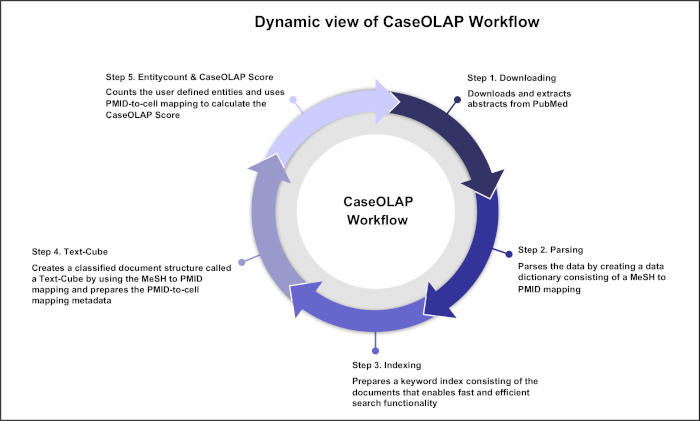
Figure 1. Dynamic view of the CaseOLAP Workflow. This figure represents the 5 major steps in the CaseOLAP workflow. In step 1, the workflow begins by downloading and extracting textual documents (e.g., from PubMed). In step 2, extracted data are parsed to create a data dictionary for each document as well as a MeSH to PMID mapping. In step 3, data indexing is conducted to facilitate fast and efficient entity search. In step 4, implementation of user-provided category information (e.g., root MeSH for each cell) is carried out to construct a Text-Cube. In step 5, the entity count operation is implemented over index data to calculate the CaseOLAP scores. These steps are repeated in an iterative manner to update the system with the latest information available in a public database (e.g., PubMed). Please click here to view a larger version of this figure.
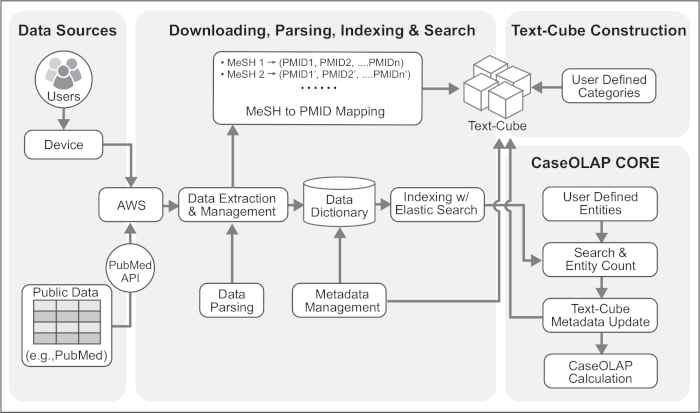
Figure 2. Technical Architecture of the CaseOLAP Workflow. This figure represents the technical details of the CaseOLAP workflow. Data from the PubMed repository are obtained from the PubMed FTP server. The user connects to the cloud server (e.g., AWS connectivity) via their device and creates a Download Pipeline which downloads and extracts the data to a local repository in the cloud. Extracted data are structured, verified, and brought to a proper format with a Data Parsing Pipeline. Simultaneously, a MeSH to PMID mapping table is created during the parsing step, which is used for Text-Cube construction. Parsed data are stored as a JSON like key-value dictionary format with document metadata (e.g., PMID, MeSH, publishing year). The Indexing step further improves the data by implementing Elasticsearch to handle bulk data. Next, the Text-Cube is created with user-defined categories by implementing MeSH to PMID mapping. When the Text-Cube formation and Indexing steps are completed, an entity count is conducted. Entity count data are implemented to the Text-Cube metadata. Finally, the CaseOLAP score is calculated based on the underlying Text-Cube structure. Please click here to view a larger version of this figure.

Figure 3. A sample of a parsed document. A sample of parsed data is presented in this figure. The parsed data are arranged as a key-value pair which is compatible with indexing and document metadata creation. In this figure, a PMID (e.g., "25896987") is serving as a key and collection of associated information (e.g., Title, Journal, Publishing date, Abstract, MeSH, Substances, Department and Location) are as value. The very first application of such document metadata is the construction of MeSH to PMID mapping (Figure 5 and Table 2), which is later implemented to create the Text-Cube and to calculate the CaseOLAP score with user-provided entities and categories. Please click here to view a larger version of this figure.
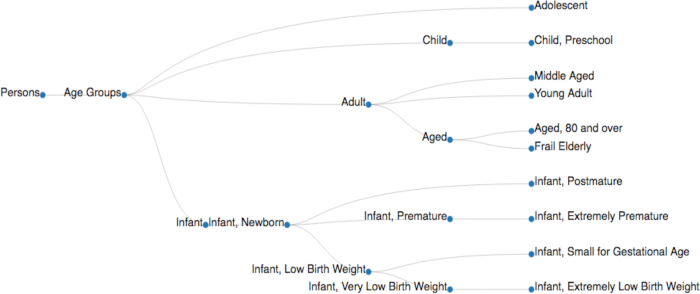
Figure 4. A sample of a MeSH tree. The 'Age Groups' MeSH tree is adapted from the tree data structure available in the NIH database (MeSH Tree 2018, <https://meshb.nlm.nih.gov/treeView>). MeSH descriptors are implemented with their node IDs (e.g., Persons [M01], Age Groups [M01.060], Adolescent [M01.060.057], Adult [M01.060.116], Child [M01.060.406], Infant [M01.060.703]) to collect the documents relevant to a specific MeSH descriptor (Table 3A). Please click here to view a larger version of this figure.
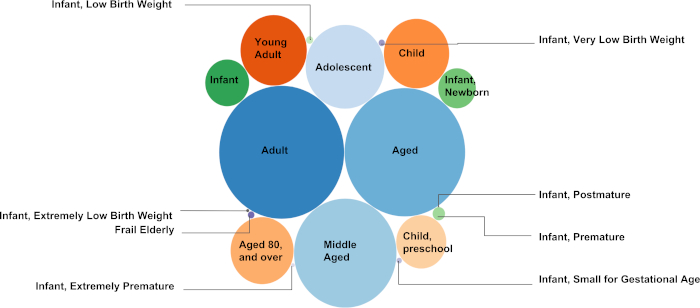
Figure 5. MeSH to PMID mapping in Age Groups. This figure presents the number of text documents (each linked with a PMID) collected under the MeSH descriptors in "Age Groups" as a bubble plot. The MeSH to PMID mapping is generated to provide the exact number of documents collected under the MeSH descriptors. A total number of 3,062,143 unique documents were collected under the 18 descendent MeSH descriptors (see Table 2). The higher the number of PMIDs selected under a specific MeSH descriptor, the larger the radius of the bubble representing the MeSH descriptor. For instance, the highest number of documents were collected under the MeSH descriptor "Adult" (1,786,371 documents), whereas the fewest number of text documents were collected under the MeSH descriptor "Infant, Postmature" (62 documents).
An additional example of MeSH to PMID mapping is given for "Nutritional and Metabolic Diseases" (https://caseolap.github.io/mesh2pmid-mapping/bubble/meta.html). A total number of 422,039 unique documents were collected under the 361 descendent MeSH descriptors in "Nutritional and Metabolic Diseases". The highest number of documents were collected under the MeSH descriptor "Obesity" (77,881 documents) followed by "Diabetes Mellitus, Type 2" (61,901 documents), whereas "Glycogen Storage Disease, Type VIII" exhibited the fewest number of documents (1 document). A related table is also available online at (https://github.com/CaseOLAP/mesh2pmid-mapping/blob/master/data/diseaseall.csv). Please click here to view a larger version of this figure.
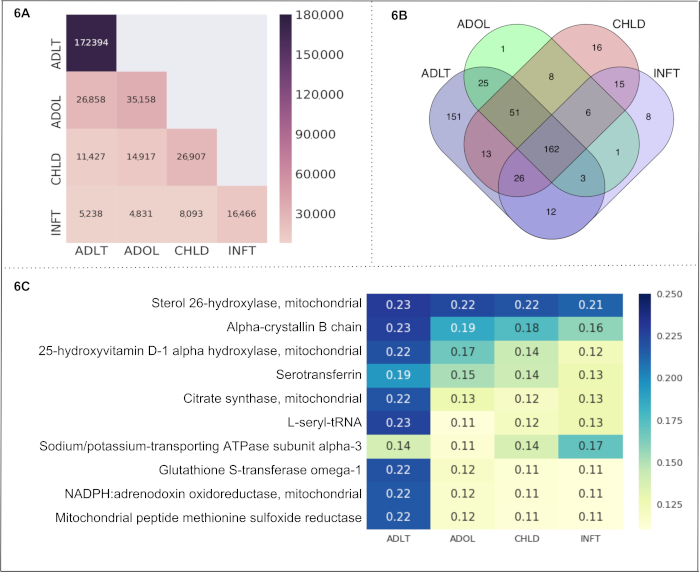
Figure 6. "Age Groups" as a use case. This figure presents the results from a use case of the CaseOLAP platform. In this instance, protein names and their abbreviations (see sample in Table 4) are implemented as entities and "Age Groups" including the cells: infant (INFT), child (CHLD), adolescent (ADOL), and adult (ADLT), are implemented as subcategories (see Table 3A). (A) Number of documents in "Age Groups": This heat map shows the number of documents distributed across the cells of "Aged Groups" (for details on the Text-Cube creation see Protocol 4 and Table 3A). A higher number of documents is presented with a darker intensity of the heatmap cell (see the scale). A single document may be included in more than one cell. The heatmap presents the number of documents within a cell along the diagonal position (e.g., ADLT contains 172,394 documents which is the highest number across all cells). The nondiagonal position represents the number of documents falling under two cells (e.g., ADLT and ADOL have 26,858 shared documents). (B). Entity count in "Age Groups": The Venn diagram represents the number of proteins found in the four cells representing "Age Groups" (INFT, CHLD, ADOL, and ADLT). The number of proteins shared within all cells is 162. The age group ADLT depicts the highest number of unique proteins (151) followed by CHLD (16), INFT (8) and ADOL (1). (C) CaseOLAP score presentation in "Age Groups": The top 10 proteins with the highest average CaseOLAP scores in each group are presented in a heat map. A higher CaseOLAP score is presented with a darker intensity of the heatmap cell (see the scale). The protein names are displayed on the left column and the cells (INFT, CHLD, ADOL, ADLT) are displayed along the x-axis. Some proteins show a strong association to a specific age group (e.g., Sterol 26-hydroxylase, alpha-crystallin B chain and L-seryl-tRNA have strong associations with ADLT, whereas Sodium/potassium-transporting ATPase subunit alpha-3 has a strong association with INFT). Please click here to view a larger version of this figure.
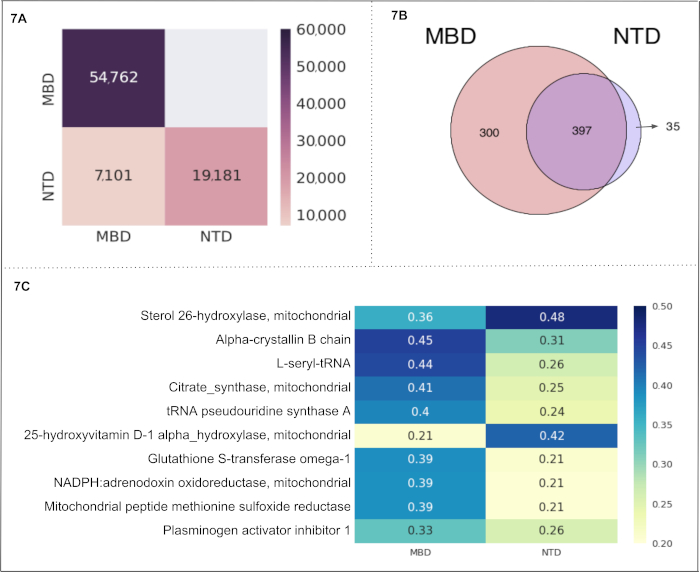
Figure 7. "Nutritional and Metabolic Diseases" as a use case: This figure presents the results from another use case of the CaseOLAP platform. In this instance, protein names and their abbreviations (see sample at Table 4) are implemented as entities and "Nutritional and Metabolic Disease" including the two cells: metabolic disease (MBD) and nutritional disorders (NTD) are implemented as subcategories (see Table 3B). (A). Number of documents in "Nutritional and Metabolic Diseases": This heatmap depicts the number of text documents in the cells of "Nutritional and Metabolic Diseases" (for details on the Text-Cube creation see Protocol 4 and Table 3B). A higher number of documents is presented with a darker intensity of the heatmap cell (see scale). A single document may be included in more than one cell. The heatmap presents the total number of documents within a cell along the diagonal position (e.g., MBD contains 54,762 documents which is the highest number across the two cells). The nondiagonal position represents the number of documents shared by the two cells (e.g., MBD and NTD have 7,101 shared documents). (B). Entity Count in "Nutritional and Metabolic Diseases": The Venn diagram represents the number of proteins found in the two cells representing "Nutritional and Metabolic Diseases" (MBD and NTD). The number of proteins shared within the two cells is 397. The MBD cell depicts 300 unique proteins, and the NTD cell depicts 35 unique proteins. (C). CaseOLAP score presentation in "Nutritional and Metabolic Diseases": The top 10 proteins with the highest average CaseOLAP scores in "Nutritional and Metabolic Diseases" are presented in a heat map. A higher CaseOLAP score is presented with a darker intensity of the heatmap cell (see scale). The protein names are displayed on the left column and cells (MBD and NTD) are displayed along the x-axis. Some proteins show a strong association to a specific disease category (e.g., alpha-crystallin B chain has a high association with metabolic disease and sterol 26-hydroxylase has a high association with nutritional disorders). Please click here to view a larger version of this figure.
| Time spent (percentage of total time) | Steps in the CaseOLAP platform | Algorithm and Data Structure of the CaseOLAP platform | Complexity of Algorithm and Data Structure | Details of the Steps |
| 40% | Downloading and Parsing |
Iteration and tree parsing algorithms | Iteration with nested loop and constant multiplication: O(n^2), O(log n). Where ‘n’ is no of iterations. | The Downloading pipeline iterates each procedure over multiple files. Parsing of a single document runs each procedure over the tree structure of raw XML data. |
| 30% | Indexing, Searching and Text Cube Creation | Iteration, Search algorithms by Elasticsearch (sorting, Lucene index, priority queues, finite state machines, bit twiddling hacks, regex queries) | Complexity related to Elasticsearch (https://www.elastic.co/) | Documents are indexed by implementing the iteration process over the data dictionary. The Text-Cube creation implements document meta-data and user-provided category information. |
| 30% | Entity Counting and CaseOLAP Calculation | Iteration in Integrity, Popularity, Distinctiveness calculation | O(1), O(n^2), multiple complexities related to caseOLAP Score calculation based on iteration types. | Entity count operation lists the documents and make an count operation over the list. The entity count data is used to calculate CaseOLAP score. |
Table 1. Algorithms and Complexities. This table presents information on the time spent (percentage of total time spent) on the procedures (e.g., downloading, parsing), data structure and details about the implemented algorithms in the CaseOLAP platform. CaseOLAP implements the professional indexing and search application called Elasticsearch. Additional information on complexities related to Elasticsearch and internal algorithms can be found at (https://www.elastic.co).
| MeSH descriptors | Number of of PMIDs collected |
| Adult | 1,786,371 |
| Middle Aged | 1,661,882 |
| Aged | 1,198,778 |
| Adolescent | 706,429 |
| Young Adult | 486,259 |
| Child | 480,218 |
| Aged, 80 and over | 453,348 |
| Child, Preschool | 285,183 |
| Infant | 218,242 |
| Infant, Newborn | 160,702 |
| Infant, Premature | 17,701 |
| Infant, Low Birth Weight | 5,707 |
| Frail Elderly | 4,811 |
| Infant, Very Low Birth Weight | 4,458 |
| Infant, Small for Gestational Age | 3,168 |
| Infant, Extremely Premature | 1,171 |
| Infant, Extremely Low Birth Weight | 1,003 |
| Infant, Postmature | 62 |
Table 2. MeSH to PMID mapping statistics. This table presents all descendant MeSH descriptors from “Age Groups” and their number of collected PMIDs (text documents). The visualization of these statistics is presented in Figure 5.
| A | Infant (INFT) | Child (CHLD) | Adolescent (ADOL) | Adult (ADLT) |
| MeSH root ID | M01.060.703 | M01.060.406 | M01.060.057 | M01.060.116 |
| Number of descendant MeSH descriptors | 9 | 2 | 1 | 6 |
| Number of PMIDs selected | 16,466 | 26,907 | 35,158 | 172,394 |
| Number of entities found | 233 | 297 | 257 | 443 |
| B | Metabolic Diseases (MBD) | Nutritional Disorders (NTD) | ||
| MeSH root ID | C18.452 | C18.654 | ||
| Number of descendant MeSH descriptors |
308 | 53 | ||
| Number of PMIDs collected | 54,762 | 19,181 | ||
| Number of entities found | 697 | 432 |
Table 3. Text-Cube Metadata. A tabular view of Text-Cube metadata is presented. The tables provide information about the categories and MeSH descriptor roots and descendants, which are implemented to collect the documents in each cell. The table also provides the statistics of the collected documents and entities. (A) “Age Groups”: This is a tabular display of “Age Groups” including infant (INFT), child (CHLD), adolescent (ADOL), and adult (ADLT) and their MeSH root IDs, number of descendant MeSH descriptors, number of selected PMIDs and number of found entities. (B) “Nutritional and Metabolic Diseases”: This is a tabular display of “Nutritional and Metabolic Diseases” including metabolic disease (MBD) and nutritional disorders (NTD) with their MeSH root IDs, number of descendant MeSH descriptors, number of selected PMIDs and the number of found entities.
| Protein names and Synonyms | Abbreviations |
| N-acetylglutamate synthase, mitochondrial, Amino-acid acetyltransferase, N-acetylglutamate synthase long form; N-acetylglutamate synthase short form; N-acetylglutamate synthase conserved domain form] | (EC 2.3.1.1) |
| Protein/nucleic acid deglycase DJ-1 (Maillard deglycase) (Oncogene DJ1) (Parkinson disease protein 7) (Parkinsonism-associated deglycase) (Protein DJ-1) | (EC 3.1.2.-) (EC 3.5.1.-) (EC 3.5.1.124)(DJ-1) |
| Pyruvate carboxylase, mitochondrial (Pyruvic carboxylase) | (EC 6.4.1.1)(PCB) |
| Bcl-2-binding component 3 (p53 up-regulated modulator of apoptosis) | (JFY-1) |
| BH3-interacting domain death agonist [BH3-interacting domain death agonist p15 (p15 BID); BH3-interacting domain death agonist p13 ; BH3-interacting domain death agonist p11 ] | (p22 BID) (BID) (p13 BID)(p11 BID) |
| ATP synthase subunit alpha, mitochondrial (ATP synthase F1 subunit alpha) | |
| Cytochrome P450 11B2, mitochondrial (Aldosterone synthase) (Aldosterone-synthesizing enzyme) (CYPXIB2) (Cytochrome P-450Aldo) (Cytochrome P-450C18) (Steroid 18-hydroxylase) | (ALDOS) (EC 1.14.15.4) (EC 1.14.15.5) |
| 60 kDa heat shock protein, mitochondrial (60 kDa chaperonin) (Chaperonin 60) (CPN60) (Heat shock protein 60) (Mitochondrial matrix protein P1) (P60 lymphocyte protein) | (HSP-60) (Hsp60) (HuCHA60)(EC 3.6.4.9) |
| Caspase-4 (ICE and Ced-3 homolog 2) (Protease TX) [Cleaved into: Caspase-4 subunit 1; Caspase-4 subunit 2] | (CASP-4) (EC 3.4.22.57)(ICH-2) (ICE(rel)-II) (Mih1) |
Table 4. Sample Entity Table. This table presents the sample of entities implemented in our two use cases: “Age Groups” and “Nutritional and Metabolic Diseases” (Figure 6 and Figure 7, Table 3A,B). The entities include protein names, synonyms, and abbreviations. Each entity (with its synonyms and abbreviations) is selected one by one and is passed through the entity search operation over indexed data (see protocol 3 and 5). The search produces a list of documents which further facilitate the entity count operation.
| Quantities | User Defined | Calculated | Equation of the quantity | Meaning of the quantity |
| Integrity | Yes | No | Integrity of user defined entities considered to be 1.0. | Represents a meaningful phrase. Numerical value is 1.0 when it is already an established phrase. |
| Popularity | No | Yes | Popularity equation in Figure 1 (Workflow and Algorithm) from reference 5, 'Materials and Methods' section. | Based on term frequency of the phrase within a cell. Normalized by total term frequency of the cell. Increase in term frequency has diminishing result. |
| Distinctiveness | No | Yes | Distinctiveness equation in Figure 1 (Workflow and Algorithm) from reference 5, 'Materials and Methods' section. | Based on term frequency and document frequency within a cell and across the neighbouring cells. Normalized by total term frequency and document frequency. Quantitatively, it is the probability that a phrase is unique in a specific cell. |
| CaseOLAP score | No | Yes | CaseOLAP score equation in Figure 1 (Workflow and Algorithm) from reference 5, 'Materials and Methods' section. | Based on Integrity, Popularity, and Distinctiveness. Numerical value always falls within 0 to 1. Quantitatively the CaseOLAP score represents the phrase-category association |
Table 5. CaseOLAP equations: The CaseOLAP algorithm was developed by Fangbo Tao and Jiawei Han et al. in 20161. Briefly, this table presents the CaseOLAP score calculation consisting of three components: integrity, popularity, and distinctiveness, and their associated mathematical meaning. In our use cases, the integrity score for proteins is 1.0 (the maximum score) because they stand as established entity names. The CaseOLAP scores in our use cases can be seen in Figure 6C and Figure 7C.
Discussion
We have demonstrated that the CaseOLAP algorithm can create a phrase based quantitative association to a knowledge-based category over large volumes of textual data for extraction of meaningful insights. Following our protocol, one can build the CaseOLAP framework to create a desired Text-Cube and quantify entity-category associations through CaseOLAP score calculation. The obtained raw CaseOLAP scores can be taken to integrative analyses including dimensionality reduction, clustering, temporal and geographical analysis, as well as the creation of a graphical database which enables semantic mapping of the documents.
Applicability of the algorithm. Examples of user-defined entities, other than proteins, could be a list of gene names, drugs, specific signs and symptoms including their abbreviations and synonyms. Furthermore, there are many choices for category selection to facilitate specific user-defined biomedical analyses (e.g., Anatomy [A], Discipline and Occupation [H], Phenomena and Processes [G]). In our two use cases, all scientific publications and their textual data are retrieved from the MEDLINE database using PubMed as the search engine, both managed by the National Library of Medicine. However, the CaseOLAP platform may be applied to other databases of interest containing biomedical documents with textual data such as the FDA Adverse Event Reporting System (FAERS). This is an open database containing information on medical adverse events and medication error reports submitted to FDA. In contrast to MEDLINE and FAERS, databases in hospitals containing electronic health records from patients are not open to the public and are restricted by the Health Insurance Portability and Accountability Act known as HIPAA.
CaseOLAP algorithm has been successfully applied to the different types of data (e.g., news articles)1. The implementation of this algorithm in biomedical documents has been made in 20185. The requirements for applicability of CaseOLAP algorithm is that each of the documents should be assigned with keywords associated with the concepts (e.g., MeSH descriptors in biomedical publications, keywords in news articles). If keywords are not found, one can apply Autophrase6,7 to collect top representative phrases and build the entity list before implementing our protocol. Our protocol does not provide the step to perform Autophrase.
Comparison with other algorithms. The concept of using a Data-Cube8,9,10 and a Text-Cube2,3,4 has been evolving since 2005 with new advancements to make data mining more applicable. The concept of Online Analytical Processing (OLAP)11,12,13,14,15 in data mining and business intelligence goes back to 1993. OLAP, in general, aggregates the information from multiple systems, and stores it in a multi-dimensional format. There are different types of OLAP systems implemented in data mining. For example (1) Hybrid Transaction/Analytical Processing (HTAP)16,17, (2) Multidimensional OLAP (MOLAP)18,19-Cube based, and (3) Relational OLAP (ROLAP)20.
Specifically, the CaseOLAP algorithm has been compared with numerous existing algorithms, specifically, with their phrase segmentation enhancements, including TF-IDF+Seg, MCX+Seg, MCX, and SegPhrase. Moreover, RepPhrase (RP, also known as SegPhrase+) has been compared with its own ablation variations, including (1) RP without the Integrity measure incorporated (RP No INT), (2) RP without the Popularity measure incorporated (RP No POP), and (3) RP without the Distinctiveness measure incorporated (RP No DIS). The benchmark results are shown in the study by Fangbo Tao et al.1.
There are still challenges on data mining which can add additional functionality over saving and retrieving the data from the database. Context-aware semantic Analytical Processing (CaseOLAP) systematically implements the Elasticsearch to build an indexing database of millions of documents (Protocol 5). The Text-Cube is a document structure built over the indexed data with user-provided categories (Protocol 6). This enhances the functionality to the documents within and across the cell of the Text-Cube and allow us to calculate term frequency of the entities over a document and document frequency over a specific cell (Protocol 8). The final CaseOLAP score utilizes these frequency calculations to output a final score (Protocol 9). In 2018, we implemented this algorithm to study ECM proteins and six heart diseases to analyze protein-disease associations. The details of this study can be found in the study by Liem, D.A. et al.5. indicating that CaseOLAP could be widely used in the biomedical community exploring a variety of diseases and mechanisms.
Limitations of the algorithm. Phrase mining itself is a technique to manage and retrieve important concepts from textual data. While discovering entity-category association as a mathematical quantity (vector), this technique is unable to figure out the polarity (e.g., positive or negative inclination) of the association. One can build the quantitative summarization of the data utilizing the Text-Cude document structure with assigned entities and categories, but a qualitative concept with microscopic granularities cannot be reached. Some concepts are continuously evolving from past till now. The summarization presented for a specific entity-category association includes all incidences throughout the literature. This may lack the temporal propagation of the innovation. In the future, we plan to address these limitations.
Future Applications. About 90% of the accumulated data in the world is in the unstructured text data. Finding a representative phrase and relation to the entities embedded in the text is a very important task for the implementation of new technologies (e.g., Machine Learning, Information Extraction, Artificial Intelligence). To make the text-data machine readable, data need to be organized in the database over which the next layer of tools could be implemented. In the future, this algorithm can be a crucial step in making data mining more functional for the retrieval of information and the quantification of the entity-category associations.
Açıklamalar
The authors have nothing to disclose.
Acknowledgements
This work was supported in part by National Heart, Lung, and Blood Institute: R35 HL135772 (to P. Ping); National Institute of General Medical Sciences: U54 GM114833 (to P. Ping, K. Watson, and W. Wang); U54 GM114838 (to J. Han); a gift from the Hellen & Larry Hoag Foundation and Dr. S. Setty; and the T.C. Laubisch endowment at UCLA (to P. Ping).
Referanslar
- Tao, F., Zhuang, H., et al. Phrase-Based Summarization in Text Cubes. IEEE Data Engineering Bulletin. , 74-84 (2016).
- Ding, B., Zhao, B., Lin, C. X., Han, J., Zhai, C. TopCells: Keyword-based search of top-k aggregated documents in text cube. IEEE 26th International Conference on Data Engineering (ICDE). , 381-384 (2010).
- Ding, B., et al. Efficient Keyword-Based Search for Top-K Cells in Text Cube. IEEE Transactions on Knowledge and Data Engineering. 23 (12), 1795-1810 (2011).
- Liu, X., et al. A Text Cube Approach to Human, Social and Cultural Behavior in the Twitter Stream.Social Computing, Behavioral-Cultural Modeling and Prediction. Lecture Notes in Computer Science. 7812, (2013).
- Liem, D. A., et al. Phrase Mining of Textual Data to analyze extracellular matrix protein patterns across cardiovascular disease. American Journal of Physiology-Heart and Circulatory. , (2018).
- Shang, J., et al. Automated Phrase Mining from Massive Text Corpora. IEEE Transactions on Knowledge and Data Engineering. 30 (10), 1825-1837 (2018).
- Liu, J., Shang, J., Wang, C., Ren, X., Han, J. Mining Quality Phrases from Massive Text Corpora. Proceedings ACM-Sigmod International Conference on Management of Data. , 1729-1744 (2015).
- Lee, S., Kim, N., Kim, J. A Multi-dimensional Analysis and Data Cube for Unstructured Text and Social Media. IEEE Fourth International Conference on Big Data and Cloud Computing. , 761-764 (2014).
- Lin, C. X., Ding, B., Han, J., Zhu, F., Zhao, B. Text Cube: Computing IR Measures for Multidimensional Text Database Analysis. IEEE Data Mining. , 905-910 (2008).
- Hsu, W. J., Lu, Y., Lee, Z. Q. Accelerating Topic Exploration of Multi-Dimensional Documents Parallel and Distributed Processing Symposium Workshops (IPDPSW). IEEE International. , 1520-1527 (2017).
- Chaudhuri, S., Dayal, U. An overview of data warehousing and OLAP technology. SIGMOD Record. 26 (1), 65-74 (1997).
- Ravat, F., Teste, O., Tournier, R. Olap aggregation function for textual data warehouse. ICEIS – 9th International Conference on Enterprise Information Systems, Proceedings. , 151-156 (2007).
- Ho, C. T., Agrawal, R., Megiddo, N., Srikant, R. Range Queries in OLAP Data Cubes. SIGMOD Conference. , (1997).
- Saxena, V., Pratap, A. Olap Cube Representation for Object- Oriented Database. International Journal of Software Engineering & Applications. 3 (2), (2012).
- Maniatis, A. S., Vassiliadis, P., Skiadopoulos, S., Vassiliou, Y. Advanced visualization for OLAP. DOLAP. , (2003).
- Bog, A. . Benchmarking Transaction and Analytical Processing Systems: The Creation of a Mixed Workload Benchmark and its Application. , 7-13 (2013).
- Özcan, F., Tian, Y., Tözün, P. Hybrid Transactional/Analytical Processing: A Survey. In Proceedings of the ACM International Conference on Management of Data (SIGMOD). , 1771-1775 (2017).
- Hasan, K. M. A., Tsuji, T., Higuchi, K. An Efficient Implementation for MOLAP Basic Data Structure and Its Evaluation. International Conference on Database Systems for Advanced Applications. , 288-299 (2007).
- Nantajeewarawat, E. Advances in Databases: Concepts, Systems and Applications. DASFAA 2007. Lecture Notes in Computer Science. 4443, (2007).
- Shimada, T., Tsuji, T., Higuchi, K. A storage scheme for multidimensional data alleviating dimension dependency. Third International Conference on Digital Information Management. , 662-668 (2007).

