Detection of Human Immunodeficiency Virus Type 1 (HIV-1) Antisense Protein (ASP) RNA Transcripts in Patients by Strand-Specific RT-PCR
Summary
RNA hairpins and loops can function as primers for reverse transcription (RT) in absence of sequence-specific primers, interfering with the study of overlapping antisense transcripts. We have developed a technique able to identify strand-specific RNA, and we have used it to study HIV-1 antisense protein ASP.
Abstract
In retroviruses, antisense transcription has been described in both human immunodeficiency virus type 1 (HIV-1) and human T-lymphotropic virus 1 (HTLV-1). In HIV-1, the antisense protein ASP gene is located on the negative strand of env, in the reading frame -2, spanning the junction gp120/gp41. In the sense orientation, the 3' end of the ASP open reading frame overlaps with gp120 hypervariable regions V4 and V5. The study of ASP RNA has been thwarted by a phenomenon known as RT-self-priming, whereby RNA secondary structures have the ability to prime RT in absence of the specific primer, generating non-specific cDNAs. The combined use of high RNA denaturation with biotinylated reverse primers in the RT reaction, together with affinity purification of the cDNA onto streptavidin-coated magnetic beads, has allowed us to selectively amplify ASP RNA in CD4+ T cells derived from individuals infected with HIV-1. Our method is relatively low-cost, simple to perform, highly reliable, and easily reproducible. In this respect, it represents a powerful tool for the study of antisense transcription not only in HIV-1 but also in other biological systems.
Introduction
The antisense protein (ASP) gene is an open reading frame (ORF) located on the negative strand of the human immunodeficiency virus type 1 (HIV-1) envelope (env) gene, spanning the junction gp120/gp411. Over the past 30 years, several reports have shown that the HIV ASP gene is indeed transcribed and translated2,3,4,5,6,7,8,9. Although ASP antisense transcripts have been fully characterized in vitro, until recently information about the actual production of ASP RNA in patients was still missing.
The sequence of ASP is reverse and complementary to env. This represents a major obstacle when trying to detect transcripts for ASP. Standard reverse transcription-polymerase chain reaction (RT-PCR) methods use gene-specific antisense primers to synthesize complementary DNAs (cDNAs) of the right polarity. This approach, however, does not allow to determine the orientation (sense or antisense) of the initial RNA template, since RNA hairpins or loops can prime RT in both directions in absence of primers10, a phenomenon known as RT self-priming. Most ASP investigators sidestep the problem of RT self-priming using primers tagged with sequences that are not related to HIV-111,12. This strategy, however, does not eliminate the occurrence of the phenomenon, and may lead to potential carry-over of non-specific cDNAs into the PCR11.
We have recently developed a novel strand-specific RT-PCR assay for the study of antisense RNA and we have used it for ASP RNA detection in a cohort of six HIV-infected patients, as shown in Table 1. The procedure described below has been previously published by Antonio Mancarella et al.13. In our protocol, we avoid the production of non-specific cDNAs by a dual approach. Firstly, we eliminate RNA secondary structures by denaturing RNA at high temperature (94 °C), and secondly, we reverse transcribe ASP RNA using a biotinylated ASP-specific primer and affinity-purify the resulting cDNA. By this approach, we are able to amplify only our target cDNA, since other non-specific RT products are either prevented from being generated (high temperature denaturation of RNA) or eliminated prior to PCR (affinity purification).
Protocol
This study was approved by the Institutional Review Board of the Centre Hospitalier Universitaire Vaudois (CHUV).
1. Infection of peripheral blood mononuclear cells (PBMCs) with HIV-1HXB2 strain
- Day 1: PBMCs STIMULATION
- Isolate PBMCs from a healthy donor buffy coat.
- Count and resuspend PBMCs at a concentration of 1×106/mL in complete Roswell Park Memorial Institute (RPMI) 1640 medium containing 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin (R-10), with addition of 50 U/mL of interleukin-2 (IL-2) and 3 µg/mL phytohaemagglutinin (PHA).
- Pipet up and down and split the cells in two six-well plates (3 mL of cell suspension/well).
- Incubate for 3 days at 37 °C and 5% CO2.
- Day 3: PBMCs INFECTION
NOTE: Use one of the two plates as in step 1.1.3 as negative control (do not infect these cells).
CAUTION: Perform the entire procedure in a biosafety level III laboratory (P3).- Pool PBMCs from one plate into a 50 mL falcon tube and centrifuge at 300 x g for 10 min.
- Discard the supernatant and resuspend the cells in 2.2 mL of R-10 containing 20 U/mL of IL-2 and 2 µg/mL polybrene.
- Thaw vials containing the HIV-1HXB2 virus and add 1.8 mL of viral suspension (0.376 ng/µL) to the cells.
- Incubate the cells for 2 h at 37 °C, gently swirling the tube every 30 min.
- Centrifuge the cells at 300 x g for 10 min.
- Discard the supernatant and resuspend the cells at 1×106/mL in R-10 containing IL-2 (50 U/mL).
- Transfer the cell suspension to a 24-well plate at a concentration of 1 mL/well and incubate for 5 days at 37 °C and 5% CO2.
- Harvest 1 well each day and transfer the cells to 1.5 mL tube (labeled with the day of infection).
NOTE: Before centrifugation, transfer 150 µL of cell suspension to a flow cytometry tube for PBMCs infection quality control (see step 1.3) - Centrifuge the tubes at 400 x g for 10 min and discard the supernatant.
- Freeze the cell pellets at -80 °C until use.
- PBMCs infection: quality control
- For each day of infection, collect 150 µL of infected PBMCs and transfer them into a flow cytometry tube.
- Add 1 mL of phosphate-buffered saline (PBS) and centrifuge at 400 x g for 5 min.
- Discard the supernatant and add 50 µL of PBS containing 1 µL of Aqua live/dead dye (previously diluted 1:10 with PBS).
- Incubate at 4 °C for 15 min.
- Add 1 mL of PBS, centrifuge at 400 x g for 5 min, and discard supernatant.
- Add 250 µL of Fixation/Permeabilization solution and incubate for 20 min at room temperature in the dark.
- Add 1 mL of 1x Perm/Wash Buffer (10x stock solution containing both FBS and saponin diluted 1:10) and centrifuge at 400 x g for 5 min.
- Discard the supernatant and add 50 µL of PBS containing HIV Gag p24 fluorescein isothiocyanate (FITC)-conjugated antibody (diluted 1:10).
- Incubate for 20 min at room temperature in the dark.
- Add 1 mL of PBS, centrifuge at 400 x g for 5 min, and discard the supernatant.
- Add 150 µL of x CellFIX (10x stock solution containing 10% formaldehyde, 3.55% methanol, 0.93% sodium azide diluted 1:10) and analyze cells by flow cytometry.
2. Stimulation of human CD4+ T cells with anti-CD3/CD28 antibodies
- Isolate CD4+ T cells from PBMCs of both HIV-1 infected patients and healthy donors.
- Prepare human anti-CD3/CD28 antibody mix by diluting anti-CD3 (1:100) and anti-CD28 (1:50) in PBS.
- Coat a 48-well plate by adding 200 µL of antibody mix per well to an appropriate number of wells and incubate at 37 °C for 2 h.
- Aspirate the antibody solution and add 1 mL of CD4+T cells at 1×106/mL to the anti-CD3/CD28 antibody-coated wells.
NOTE: Stimulate CD4+ T cells up to 5 days.
3. Reverse transcription
NOTE: To obtain patient-specific primers, proviral DNA isolated from CD4+T cells from each patient was amplified using HIV-1HXB2 Pan ASP primers. Patient specific primers were designed using the proviral sequence internal to the Pan ASP primers. All primers and probes used in this study are listed in Table 2.
- Isolating total RNA from cells
- Quantify RNA and eliminate DNA contamination by treating samples with DNase.
- Perform RT reactions in a volume of 50 µL. Transfer 0.1-1 µg of total RNA into an appropriate number of wells of a 96-well PCR plate. Prepare the RNA mixture by adding the following components to the RNA:
5 µL of biotinylated ASP reverse primer (20 µM)
2.5 µL of deoxynucleotide triphosphates (dNTPs) (10 mM)
25 µL of diethylpyrocarbonate (DEPC)-treated distilled water (dH2O).
Prepare the endogenous RT controls by adding 5 µL of nuclease-free water in place of the biotinylated ASP reverse primer. - Place the 96 well PCR plate into a thermocycler and heat the RNA mixture to 94 °C for 5 min.
- Immediately cool down the RNA mixture in iced water for at least 1 min.
- Prepare the reaction mixture by adding the following components to a 1.5 mL tube (calculate the total number of samples plus 1):
10 µL of 5x RT buffer
2.5 µL of 0.1 M of 1,4-dithiothreitol (DTT)
2.5 µL of RNase out (40 units/µL)
2.5 µL of RT enzyme (200 units/µL) - Transfer 17.5 µL of this mixture to each of the RNA-containing wells. Prepare the RT-minus controls replacing reverse transcriptase with 2.5 µL of nuclease-free water.
- Mix gently and incubate the plate at 55 °C for 60 min using a thermocycler.
- Inactivate the reactions by heating at 70 °C for 15 min.
4. Affinity purification of ASP biotinylated cDNA
NOTE: Do not purify RT-minus reactions.
- Prepare 1 L of 2x washing/binding buffer containing 10 mM Tris-HCl (pH 7.5), 1 mM ethylenediaminetetraacetic acid (EDTA) and 2 M NaCl. Filter the solution.
- Prepare 50 mL of 1x washing/binding buffer using PCR grade water.
- For each reaction, use 10 µL of streptavidin-conjugated magnetic beads. Based on the number of reactions, transfer an appropriate volume of beads to a 1.5 mL tube.
- Wash the beads by adding an equal volume of 2x washing/binding buffer and vortex for 15 s.
- Place the tube into a magnetic separation rack and incubate for 3 min at room temperature.
- Carefully remove the supernatant without disturbing the beads and add the same volume of washing/binding 2x buffer as the initial volume of beads.
- Vortex for 30 s and transfer 10 µL of the beads suspension to an appropriate number of 1.5 mL tubes.
- Place the tubes in a magnetic separation rack and incubate for 3 min at room temperature.
- Discard the supernatant and add 50 µL of washing/binding 2x buffer.
- Add 50 µL of biotinylated cDNA to the corresponding tubes containing the beads.
- Incubate for 30 min at room temperature rotating gently.
- Separate the beads from the supernatant by a magnet separation rack for 3 min at room temperature.
- Wash the beads by adding 200 µL of 1x washing/binding buffer. Place the tubes in a magnet separation rack for 3 min at room temperature and discard the supernatant. Repeat twice.
- Resuspend the beads in 10 µL of PCR grade water.
5. Standard PCR
NOTE: The aim of the standard PCR is to amplify the entire ORF of the ASP gene. The amplification products are then cloned into pCR2.1 plasmid to develop standard curves for ASP RNA quantification by real-time PCR (see paragraph real time PCR).
- Perform standard PCRs in a total volume of 50 µL. Prepare an appropriate volume of PCR master mix (number of samples plus 1). For each sample, add the following components to a 1.5 mL tube:
1 µL of 10 µM ASP primers (forward and reverse)
1 µL of 10 mM dNTPs
Magnesium chloride (MgCl2) (concentration depends on the primers used)
dH2O to a final volume of 49 µL- Mix well, quickly spin down contents and transfer 49 µL/well of the PCR master mix to a 96-well PCR plate.
- Carefully vortex the tubes containing the beads used for ASP cDNA affinity purification for 15 s.
- Add 1 µL of beads into the corresponding wells and run the PCR using the following program: 95 °C for 2 min, 40 cycles of 95 °C for 30 s, 55 °C for 30 s, and 68 °C for 40 s, then 68 °C for 7 min.
- Separate the PCR products onto 1 % agarose gel, cut the bands and clone the amplified products into pCR2.1 plasmid.
- Sequence and analyze the sequences of each clone.
6. Real time quantitative PCR (qPCR)
NOTE: Develop patient-specific primers and probes using the approach described in step 3 “Reverse Transcription”. For qPCR include ASP plasmid dilutions for standard curves. Plasmids containing patient-specific inserts are developed as mentioned in the note at the beginning of step 5 “Standard PCR”.
- Prepare the standard curve using 1:10 ASP plasmid serial dilutions starting from 3 x 106 copies/µL down to 3 copies/µL.
- First round PCR (pre-amp). Perform reactions in a total volume of 25 µL. Prepare an appropriate volume of PCR master mix (number of samples plus 1). For each sample, add the following components to a 1.5 mL tube:
0.5 µL of ASP or env primer mix (final concentration 0.2 µM each) (forward and reverse)
0.5 µL of dNTPs mix (final concentration 0.2 mM each)
MgCl2 (concentration depends of the primer pairs)
dH2O to a final volume of 24 µL - Transfer 24 µL of PCR master mix to an appropriate number of wells of a 96-well PCR plate.
- Carefully vortex the tubes containing the beads with the cDNA for 15 s.
- Add 1 µL of beads to the corresponding wells. Do the same for plasmid dilutions.
- Run the PCR using the following algorithm: denaturation for 2 min at 95 °C ; 30 s at 95 °C, 30 s at 55 °C, 40 s at 68 °C for 18 cycles; extension at 68 °C for 7 min.
- Dilute the first round PCRs reactions 1:5 with PCR grade water.
- Second round qPCR. Perform reactions in a total volume of 20 µL. Prepare an appropriate volume of PCR master mix (number of samples plus 1). For each sample, add the following components to a 1.5 mL tube:
1.8 µL 10 µM primers (forward and reverse)
1 µL of 5 µM probe
6.2 µL of dH2O - Transfer 19 µL of qPCR master mix into an appropriate number of wells of a 96-well qPCR plate and add 1 µL of the diluted first round PCR reactions to the corresponding wells. Do the same for the plasmid dilutions.
- Run the qPCR with the following algorithm: denaturation for 10 min at 95 °C; 15 s at 95 °C, 1 min at 60 °C for 40 cycles.
- First round PCR (pre-amp). Perform reactions in a total volume of 25 µL. Prepare an appropriate volume of PCR master mix (number of samples plus 1). For each sample, add the following components to a 1.5 mL tube:
Representative Results
High temperature RNA denaturation coupled to affinity purification of biotinylated cDNA prevents amplification of non-specific ASP products during PCR in PBMCs infected in vitro and in CD4+ T cells isolated from patients. RT self-priming has been shown to occur during reverse transcription of antisense RNAs10,14,15,16,17. In order to prevent this phenomenon, we developed a novel approach using the technique originally described by Heist et al.10. Our procedure involves high temperature denaturation of the RNA prior to RT and the use of biotinylated primers to perform RT, followed by affinity-purification of the biotinylated cDNAs onto streptavidin-coated magnetic beads13. The cDNAs obtained by this method can be used for downstream applications including standard PCR or qPCR.
The experimental conditions for antisense transcription were first tested in PBMCs from one healthy donor infected in vitro with HIV-1HXB2, the reference sequence for HIV-1 Subtype B (all our patients are infected with isolates of this subtype). Table 2 shows the sequences of primers and probes used in this study. Our initial RT reactions were done using a regular, non-biotinylated antisense primer (ASP R1) and resulted in the successful amplification of ASP (ASP F1-R1 primer pair) (Figure 1A, Lanes 1 – 3). However, by this approach, we also amplified a band of the same molecular weight in primer-minus RT reactions controls (Lanes 4 – 6). The complete lack of signal in our negative controls (Lanes 7 – 8) indicated that the non-specific template was coming from the RT reaction and not from cross-contamination of samples.
To bypass the problem created by RT self-priming, we decided to use a method previously described by Haist et al.10, in which the specific antisense primer is labelled with biotin so that the resulting biotinylated cDNA corresponding to the antisense orientation can be purified prior to PCR and selectively amplified. By this approach, we were able to amplify the ASP sequence (Figure 1B, Lanes 1 – 3) with greatly reduced contamination from the non-specific cDNA in the primer-minus controls (Figure 1B, Lanes 4-6).
Optimization of this method was achieved by complete denaturation of the RNA prior to RT at 94 °C, followed by immediate cooling onto iced water. As shown in Figure 2A,B, the ASP band is effectively amplified, whereas the non-specific products have disappeared.
ASP RNA is detected in CD4+ T cells from patients with detectable viraemia and in absence of therapy, following stimulation with anti-CD3/CD28. ASP RNA was undetectable in either unfractionated PBMCs or unstimulated CD4+ T cells isolated from HIV patients (data not shown). However, it could be easily detected in CD4 cells isolated from three HIV-positive subjects, MP135, MP140, and MP148 (Table 1), following stimulation with anti-CD3/CD28. The kinetics of ASP RNA expression measured by qPCR in these three patients are shown in Figure 3.
CD4 cells isolated from patients undergoing ART and with non-detectable viraemia, may produce small amounts of ASP RNA when stimulated with anti-CD3/CD28. Quantification of ASP RNA levels in two of these patients (MP071, MP146) by qPCR shows that in these conditions ASP RNA is detected in low levels (10-15 copies/million CD4+ T cells) at 3-5 days post-stimulation (Figure 4). In one patient however (MP069), no ASP RNA could be detected at any time point (data not shown).
ASP and env RNAs have similar patterns of expression in one untreated patient with detectable viraemia (MP140). Two patients were analyzed, one untreated (MP140) and one treated (MP146). In MP140 we could detect both ASP and env. Although their levels of transcription were dissimilar (Figure 5A), the expression curve of these two genes over time was identical, which can be visualized plotting data on a logarithmic scale (Figure 5B). In patient MP146, which was treated and whose viraemia was below the levels of detection, ASP and env were barely detectable and only after several days of stimulation (Figure 5C).
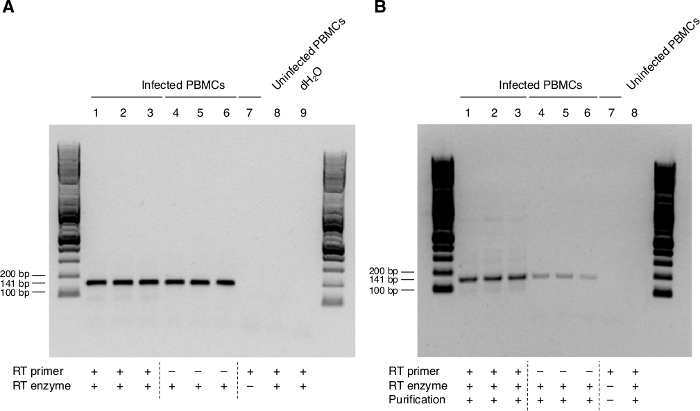
Figure 1: Expression of ASP RNA in PBMCs from one HIV-negative individual infected in vitro with HIV-1HXB2 by standard and modified RT-PCR. (A) Detection of ASP RNA using standard RT-PCR. ASP is amplified from cDNA synthesized in the presence of the non-biotinylated ASP-specific antisense primer ASP R1 (Lanes 1–3). The same band is also found in primer-minus RT controls (Lanes 4–6). (B) Visualization of ASP RNA by biotinylation of the antisense primer and cDNA purification (Lanes 1-3). A band of decreased intensity is still detected in primer-minus controls (Lanes 4–6). No bands are present in negative controls. Previously published in the article "Detection of antisense protein (ASP) RNA transcripts in individuals infected with human immunodeficiency virus type 1 (HIV-1)", by Mancarella et al.13 (J Gen Virol 100(5):863-876. doi: 10.1099/jgv.0.001244). Please click here to view a larger version of this figure.
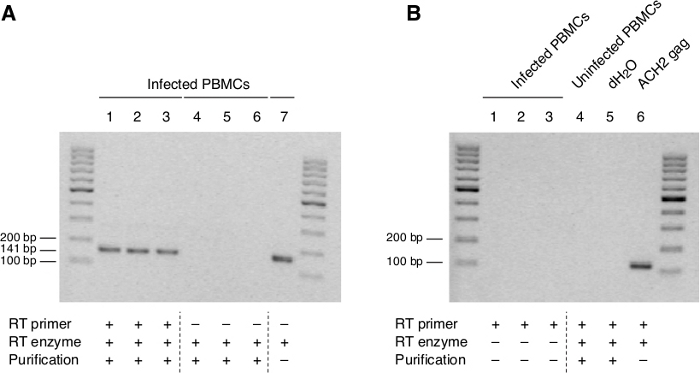
Figure 2: ASP RNA is expressed in PMBCs of one HIV-negative individual following infection with HIV-1HXB2. (A) The ASP band is easily detected RNA which has been fully denatured and reverse-transcribed using the biotinylated RT primer (ASP R1) (Lanes 1–3) but not in purified cDNA from primer-minus controls (Lanes 4–6). (B) No signal corresponding to ASP is detected in RT-minus controls of RNA from infected PBMCs, uninfected PBMCs or water, although a clear band is visible in gag positive control from ACH-2 cells. Previously published in the article "Detection of antisense protein (ASP) RNA transcripts in individuals infected with human immunodeficiency virus type 1 (HIV-1)", by Mancarella et al.13 (J Gen Virol 100(5):863-876. doi: 10.1099/jgv.0.001244). Please click here to view a larger version of this figure.
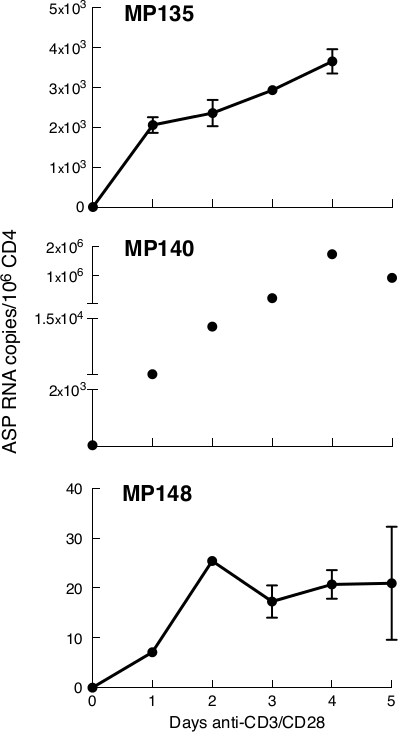
Figure 3: In anti-CD3/CD28-stimulated CD4+ T cells isolated from three untreated patients, ASP RNA production is readily detectable and peaks between day 2 and 4 post-stimulation. In MP135 and MP140, expression of ASP peaked at day 4 post-stimulation, while in MP148 it peaked at day 2. ASP levels are expressed as RNA copies/million CD4+ T cells. Points in the time course correspond to the mean value of triplicate PCR reactions. Previously published in the article "Detection of antisense protein (ASP) RNA transcripts in individuals infected with human immunodeficiency virus type 1 (HIV-1)", by Mancarella et al.13 (J Gen Virol 100(5):863-876. doi: 10.1099/jgv.0.001244). Please click here to view a larger version of this figure.
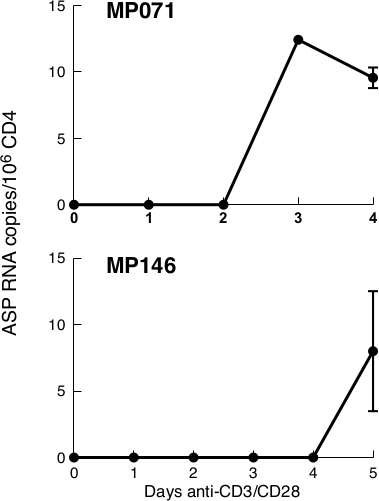
Figure 4: In two aviremic patients undergoing ART, ASP is barely detectable only after a few days of stimulation. In both our patients, ASP could not be detected at day 0. In Patient MP071, low levels of ASP could be detected at day 3, whereas in Patient MP146, we had to wait until day 5 in order to see some levels of RNA. ASP levels are expressed as RNA copies/million CD4+ T cells. Points in the time course correspond to the mean value of triplicate PCR reactions. Previously published in the article "Detection of antisense protein (ASP) RNA transcripts in individuals infected with human immunodeficiency virus type 1 (HIV-1)", by Mancarella et al.13 (J Gen Virol 100(5):863-876. doi: 10.1099/jgv.0.001244). Please click here to view a larger version of this figure.
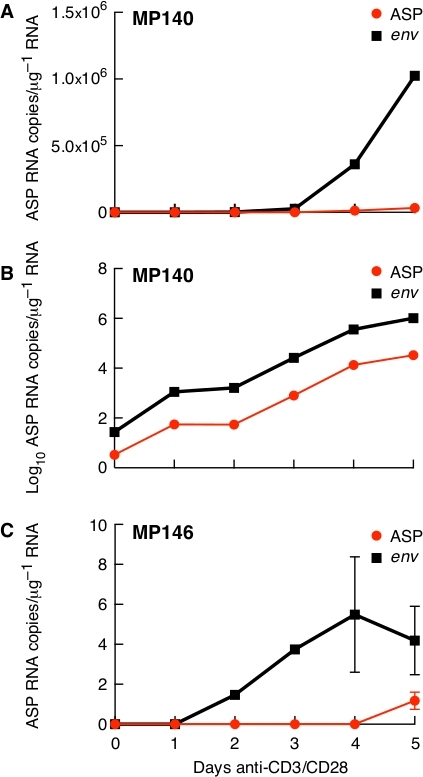
Figure 5: Expression of ASP and env in anti-CD3/CD28-stimulated CD4+ T cells in treated (MP146) and untreated (MP140) patients. (A) In the untreated patient (MP140), env is expressed at higher levels than ASP; (B) The same data plotted on a logarithmic scale show that ASP and env expression are characterized by a similar profile over-time; (C) Expression kinetics of ASP and env in one patient with undetectable viraemia and undergoing ART (MP146). Previously published in the article "Detection of antisense protein (ASP) RNA transcripts in individuals infected with human immunodeficiency virus type 1 (HIV-1)", by Mancarella et al.13 (J Gen Virol 100(5):863-876. doi: 10.1099/jgv.0.001244). Please click here to view a larger version of this figure.
| Patient ID | Age | Sex | Stage of HIV infection | Clade | Viral load (copies/ml) | CD4 Count (cells/µl) | ART status |
| MP135 | 44 | M | C3 | B | 1.6×105 | 176 | Untreated |
| MP140 | 23 | M | A2 | B | 3.6×104 | 427 | Untreated |
| MP148 | 37 | M | A1 | B | 2.0×104 | 717 | Untreated |
| MP069 | 42 | M | A1 | B | <20 | 1309 | Treated |
| MP071 | 47 | M | C3 | B | <20 | 167 | Treated |
| MP146 | 59 | M | C3 | B | <20 | 385 | Treated |
Table 1: Patients' features. Previously published in the article "Detection of antisense protein (ASP) RNA transcripts in individuals infected with human immunodeficiency virus type 1 (HIV-1)", by Mancarella et al.13 (J Gen Virol 100(5):863-876. doi: 10.1099/jgv.0.001244).
| Primer/Probe Name | Primer sequence (5' to 3') |
| ASP F1 | TTAGGAGTAGCACCCACCAA |
| ASP R1 | GAACCCAAGGAACAAAGCTC |
| PAN ASP F | ACCAAGCCTCCTACTATCATTATG |
| PAN ASP R | GCACATTGTAACATTAGTAGAGCA |
| ASP MP135, 146, 071 F | CCCAAGAACCCAAGGAACATAG |
| ASP MP135, 146, 071 R | CATTAGGAATAGCACCCACCAA |
| ASP MP135, 146, 071 Probe FAM/TAMRA | TCTCTGCACCACTCTTCTCTTTGCC |
| ASP MP140 F | CCCATAGTGCTTCCTGCTATTC |
| ASP MP140 R | AGAAGAGTGGTGCAGAGAGA |
| ASP MP140 Probe FAM/TAMRA | AGCTCCTATTGTTCCCACTGCTCT |
| ASP MP148 F | CTCTCTGCACCACTCTTCTTT |
| ASP MP148 R | AGACCTGGAGGAGGAGATATG |
| ASP MP148 Probe FAM/TAMRA | TGGTGGGTGCTACTCCTAATGGTT |
| ENV MP140 F | AGAAGAGTGGTGCAGAGAGA |
| ENV MP140 R | CCCATAGTGCTTCCTGCTATTC |
| ENV MP140 Probe FAM/TAMRA | AGCTCCTATTGTTCCCACTGCTCT |
| ENV MP146 F | CATTAGGAATAGCACCCACCAA |
| ENV MP146 R | CCCAAGAACCCAAGGAACATAG |
| ENV MP146 Probe FAM/TAMRA | TCTCTGCACCACTCTTCTCTTTGCC |
Table 2: RT-PCR primers and probes
Discussion
In this report we describe a strand-specific RT assay applied to the detection of ASP RNA in CD4+ T cells isolated from individuals infected with HIV-1. The occurrence of non-specific priming during RT hampers the detection of RNA transcripts with the right polarity, leading to misinterpretation of the results. Previous groups have developed several strategies aimed at preventing primer-independent cDNA synthesis during the RT reaction. Tagging the reverse primer at the 3' end with sequences not related to HIV has proved effective in achieving strand-specific amplification6,8,9. However, this approach only allows for making the falsely primed cDNA undetectable rather than to avoid it, with the risk of it leaking into the PCR reaction, causing amplification of non-specific DNA products (i.e., env instead of ASP).
Our initial RT-PCR attempts at detecting ASP RNA were performed using a standard antisense primer. Amplifications were successful, as we obtained bands of the right molecular weight, but bands of the same size were also present in our primer-minus controls. Based on these results, we used a different approach, performing RT with a biotinylated specific antisense primer, as described by Haist et al.10. Although our preliminary experiments resulted in a drastic decrease of the RT self-priming, we could not fully remove non-specific products of amplification. Haist and coworkers eliminate non-specific cDNA products by washing the beads in high stringency conditions10. In our method, we removed completely the source of self-priming in the form of RNA secondary structures using high denaturation temperatures to fully linearize the RNA template.
We show that ASP RNA is expressed in anti-CD3/CD28-stimulated CD4+ T lymphocytes. Detection of ASP, however, cannot be achieved in cells in absence of stimulation. Our data are somewhat dissimilar from those by Zapata and coworkers, who have reported expression of low levels of ASP in resting CD4+ cells isolated from patients undergoing ART9. Based on these results, they propose that ASP RNA may play a role in regulating HIV latency. Our finding that in the same individual the kinetics of expression of ASP and env are characterized by the same profile is not consistent with the Zapata's model9. In fact, if really ASP was involved in the regulation of latency, its expression profile should be opposite to the one observed for env18.
We also observed that env transcription levels are over 2 logs higher than those of ASP, at least in patients in absence of therapy. These data are in agreement with studies by Laverdure et al.8 showing that in primary CD4+ cells infected in vitro, 3' LTR antisense transcription is much lower (up to 1000-fold) than 5' sense transcription. Our results indicate that ASP is expressed in infected CD4 lymphocytes regardless of the stage of the disease as long as cells are properly stimulated. In addition our data demonstrate that ASP is expressed in cells from patients undergoing ART treatment, although the level of expression in these cells is lower than in cells from patients in absence of therapy.
In summary, we suggest a reliable strand-specific RT assay aimed at preventing primer-independent cDNA synthesis. ASP and env are two genes which overlap to each other in opposite orientation, a feature that makes their study very challenging. Our approach, which allows for selectively capturing cDNAs retrotranscribed from RNA with both negative and positive polarity, represents an optimal and effective tool for the study of these two genes in particular, and genes overlapping in their antisense orientation in general.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank Patrizia Amelio, Alessandra Noto, Craig Fenwick, and Matthieu Perreau for always being available to discuss our work and all the people in the Laboratory of AIDS Immunopathogenesis for their precious technical assistance. We also would like to thank John and Aaron Weddle from VSB Associated Inc. who contributed with excellent artwork. Finally, many special thanks to all the Patients, without whom this work would not have been possible. This work received no specific grant from any funding agency.
Materials
| BD LSR II | Becton Dickinson | ||
| BigDye Terminator v1.1 Cycle Sequencing Kit | Applied Biosystem, Thermo Fisher Scientific | 4337450 | |
| dNTP Set (100 mM) | Invitrogen, Thermo Fisher Scientific | 10297018 | |
| Dynabeads M-280 Streptavidin | Invitrogen, Thermo Fisher Scientific | 11205D | |
| EasySep Human CD4+ T Cell Isolation Kit | Stemcell Technologies | 19052 | |
| Fetal Bovine Serum | Biowest | S1010-500 | |
| Fixation/Permeabilization Solution Kit | Becton Dickinson | 554714 | |
| HIV Gag p24 flow cytometry antibody – Kc57-FITC | Beckman Coulter | 6604665 | |
| Human IL-2 | Miltenyi Biotec | 130-097-743 | |
| Lectin from Phaseolus vulgaris (PHA) | Sigma-Aldrich | 61764-1MG | |
| LIVE/DEAD Fixable Yellow Dead Cell Stain Kit, for 405 nm excitation | Invitrogen, Thermo Fisher Scientific | L34967 | |
| Mouse Anti-Human CD28 | Becton Dickinson | 55725 | |
| Mouse Anti-Human CD3 | Becton Dickinson | 55329 | |
| Primers and Probes | Integrated DNA Technologies (IDT) | ||
| Penicillin-Streptomycin | BioConcept | 4-01F00-H | |
| Platinum Taq DNA Polymerase High Fidelity | Invitrogen, Thermo Fisher Scientific | 11304011 | |
| Polybrene Infection / Transfection Reagent | Sigma-Aldrich | TR-1003-G | |
| RNeasy Mini Kit | Qiagen | 74104 | |
| Roswell Park Memorial Institute (RPMI) 1640 Medium | Gibco, Thermo Fisher Scientific | 11875093 | |
| StepOnePlus Real-Time PCR System | Applied Biosystem, Thermo Fisher Scientific | 4376600 | |
| SuperScript III Reverse Transcriptase | Invitrogen, Thermo Fisher Scientific | 18080044 | |
| TaqMan Gene Expression Master Mix | Applied Biosystem, Thermo Fisher Scientific | 4369016 | |
| TOPO TA Cloning Kit for Subcloning, with One Shot TOP10 chemically competent E. coli cells | Invitrogen, Thermo Fisher Scientific | K450001 | |
| TURBO DNase (2 U/µL) | Invitrogen, Thermo Fisher Scientific | AM2238 | |
| Veriti Thermal Cycler | Applied Biosystem, Thermo Fisher Scientific | 4375786 |
References
- Miller, R. H. Human immunodeficiency virus may encode a novel protein on the genomic DNA plus strand. Science. 239 (4846), 1420-1422 (1988).
- Vanheebrossollet, C., et al. A Natural Antisense Rna Derived from the Hiv-1 Env Gene Encodes a Protein Which Is Recognized by Circulating Antibodies of Hiv+ Individuals. Virology. 206 (1), 196-202 (1995).
- Briquet, S., Vaquero, C. Immunolocalization studies of an antisense protein in HIV-1-infected cells and viral particles. Virology. 292 (2), 177-184 (2002).
- Clerc, I., et al. Polarized expression of the membrane ASP protein derived from HIV-1 antisense transcription in T cells. Retrovirology. 8, 74 (2011).
- Landry, S., et al. Detection, characterization and regulation of antisense transcripts in HIV-1. Retrovirology. 4, 71 (2007).
- Kobayashi-Ishihara, M., et al. HIV-1-encoded antisense RNA suppresses viral replication for a prolonged period. Retrovirology. 9, 38 (2012).
- Barbagallo, M. S., Birch, K. E., Deacon, N. J., Mosse, J. A. Potential control of human immunodeficiency virus type 1 asp expression by alternative splicing in the upstream untranslated region. DNA Cell Biol. 31 (7), 1303-1313 (2012).
- Laverdure, S., et al. HIV-1 Antisense Transcription Is Preferentially Activated in Primary Monocyte-Derived Cells. Journal of Virology. 86 (24), 13785-13789 (2012).
- Zapata, J. C., et al. The Human Immunodeficiency Virus 1 ASP RNA promotes viral latency by recruiting the Polycomb Repressor Complex 2 and promoting nucleosome assembly. Virology. 506, 34-44 (2017).
- Haist, K., Ziegler, C., Botten, J. Strand-Specific Quantitative Reverse Transcription-Polymerase Chain Reaction Assay for Measurement of Arenavirus Genomic and Antigenomic RNAs. PLoS One. 10 (5), 0120043 (2015).
- Lerat, H., et al. Specific detection of hepatitis C virus minus strand RNA in hematopoietic cells. The Journal of Clinical Investigation. 97 (3), 845-851 (1996).
- Tuiskunen, A., et al. Self-priming of reverse transcriptase impairs strand-specific detection of dengue virus RNA. J Gen Virol. 91, 1019-1027 (2010).
- Mancarella, A., et al. Detection of antisense protein (ASP) RNA transcripts in individuals infected with human immunodeficiency virus type 1 (HIV-1). Journal of General Virology. , (2019).
- Peyrefitte, C. N., Pastorino, B., Bessaud, M., Tolou, H. J., Couissinier-Paris, P. Evidence for in vitro falsely-primed cDNAs that prevent specific detection of virus negative strand RNAs in dengue-infected cells: improvement by tagged RT-PCR. J Virol Methods. 113 (1), 19-28 (2003).
- Boncristiani, H. F., Di Prisco, G., Pettis, J. S., Hamilton, M., Chen, Y. P. Molecular approaches to the analysis of deformed wing virus replication and pathogenesis in the honey bee, Apis mellifera. Virol J. 6, 221 (2009).
- Boncristiani, H. F., Rossi, R. D., Criado, M. F., Furtado, F. M., Arruda, E. Magnetic purification of biotinylated cDNA removes false priming and ensures strand-specificity of RT-PCR for enteroviral RNAs. J Virol Methods. 161 (1), 147-153 (2009).
- Craggs, J. K., Ball, J. K., Thomson, B. J., Irving, W. L., Grabowska, A. M. Development of a strand-specific RT-PCR based assay to detect the replicative form of hepatitis C virus RNA. J Virol Methods. 94 (1-2), 111-120 (2001).
- Barbeau, B., Mesnard, J. M. Making sense out of antisense transcription in human T-cell lymphotropic viruses (HTLVs). Viruses. 3 (5), 456-468 (2011).

