The Circular Dichroism Spectroscopy Technique to Study DNA-Protein Interactions
Abstract
Source: Arya, V., et al. CD Spectroscopy to Study DNA-Protein Interactions. J. Vis. Exp. (2022).
This video demonstrates the circular dichroism (CD) spectroscopy technique to study conformational changes occurring in DNA in the presence of ATP-dependent chromatin remodeling protein. The changes in the CD spectra measured in the presence and absence of ATP hydrolysis indicate the importance of ATP in the ability of the remodeler to induce conformational changes in the bound DNA.
Protocol
1. Working concentration of the reaction components
- Prepare the working concentrations of buffers for CD freshly and keep them at 4 °C before setting up the reactions.
NOTE: For the CD reactions described in this paper, the working concentrations of components are as follows: Sodium phosphate buffer (pH 7.0) 1 mM, ATP 2 mM, DNA 500 nM, Protein 1 µM, MgCl2 10 mM, EDTA 50 mM, ADAADiN 5 µM.
2. Choosing and preparation of CD cuvettes
- Collect CD spectra in high-transparency quartz cuvettes. Use rectangular or cylindrical cuvettes.
NOTE: A CD quartz cuvette (nominal volume of 0.4 mL, path length of 1 mm) was used for all the reactions described in this paper. - Use a cuvette cleaning solution to clean the cuvette. Add 1% cuvette cleaning solution in water to make 400 µL of the solution, pour it into the cuvette, and incubate it at 37 °C for 1 h.
- Wash the cuvette with water several times to clean the cuvette. Take a scan of the water or buffer in the cuvette to check whether it is clean.
NOTE: The water or buffer must give a reading in the 0 to 1 mdeg range.
3. Preparation of proteins and DNA oligonucleotide
- Keep the volume of the protein below 50 µL in the reaction to minimize the amounts of the buffer components that sometimes cause the formation of ambiguous peaks. Keep the protein on the ice throughout the experiment to avoid any degradation.
- Use PAGE-purified DNA oligonucleotides in the reactions.
NOTE: In the reactions described here, DNA was used both in native as well as heat-cooled forms (fast-cooled (FC) and slow-cooled (SC)). Fast cooling promotes intramolecular bonding in the DNA, yielding more secondary structures. In contrast, slow cooling promotes intermolecular bonding in the DNA, resulting in fewer secondary structures. - For fast cooling, heat DNA at 94 °C for 3 min on the heating block and immediately cool it on ice. For slow-cooling, heat DNA at 94 °C for 3 min and allow it to cool to room temperature at a rate of 1 °C per minute.
4. Setting up control experiments to record the baseline spectra
- Keep the reaction volume at 300 µL in all the reactions. Set up a total of 5 baseline reactions in 1.5 mL centrifuge tubes, one by one, as follows: i) Buffer + Water; ii) Buffer + MgCl2 + ATP + Water; iii) Buffer + MgCl2 + ATP + Protein + Water; iv) iii + EDTA or ADAADiN; v) Buffer + Protein +Water.
5. Setting up the experiments to record CD spectra
- Set up a total of 5 reactions, one by one, in 1.5 mL centrifuge tubes as follows: i) Buffer + DNA + Water; ii) Buffer + DNA + MgCl2 + ATP + Water; iii) Buffer + DNA + MgCl2 + ATP + Protein + Water; iv) iii + EDTA or ADAADiN; v) Buffer + DNA + Protein +Water.
6. Recording scan
- Turn on the gas and switch on the CD spectrometer.
- Switch on the lamp after 10-15 min. Switch on the water bath and set the holder temperature at 37 °C.
- Open the CD spectrum software.
- Set the temperature to 37 °C.
- Set the wavelength range at 180 – 300 nm.
- Set the time per point to 0.5 s.
- Set the scan number to 5.
- Click on Pro-Data Viewer, make a new file, and rename it with details about the experiment and date.
- Keep all the reaction components on ice to avoid any degradation. Make the baselines and reactions, one by one, in centrifuge tubes and mix them by pipetting. Transfer the reaction mix to the cuvette carefully, ensuring that there are no air bubbles.
- If performing a time-course experiment, incubate the reactions at 37 °C for the required time and take the scan. Add EDTA to the buffer containing the DNA, ATP, Mg+2, and protein to stop ATP hydrolysis.
- Increase the concentration of EDTA and its incubation time to inhibit ATPase activity completely.
- Subtract the baselines from the corresponding reactions in the software (e.g., subtract reaction 1 from baseline 1). Smoothen the data either in the CD spectrum software or in the data plotting software. Plot the data in the data plotting software.
NOTE: Subtracting the baselines from the corresponding reactions will give the net CD spectra of only DNA.
7. Data analysis and interpretation
- Use the formula given by eq (1) to convert the values obtained in millidegrees to mean residue ellipticity.
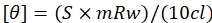
Here, S is the CD signal in millidegrees, c is the DNA concentration in mg/mL, mRw is the mean residue mass, and l is the path length in cm. - Plot a graph against wavelength and mean residue ellipticity using the data plotting software and analyze the peaks.
- To plot the graph, select the mean residue ellipticity on the Y-axis and wavelength on the X-axis and plot a straight-line graph.
NOTE: This graph will provide the characteristic peaks of different forms of DNA. The forms of DNA corresponding to the peaks can be identified using existing literature.
Disclosures
The authors have nothing to disclose.
Materials
| Adenosine 5′-triphosphate disodium salt hydrate | Sigmaaldrich | A2383 | |
| CD Quartz Cuvette | STARNA | 21-Q-1 | |
| Chirascan V100 CD spectrometer | Applied Photophysics | Not available | |
| EDTA Disodium Salt Dihydrate | SRL | 43272 | |
| Hellmanex III cleaning solution | Hellma | 9-307-011-4-507 | |
| Magnesium Chloride Hexahydrate | Fisher scientific | M33-500 | |
| Sodium Phosphate Dibasic Anhydrous | Fisher scientific | S374-500 | |
| Sodium Phosphate Monobasic Monohydrate | Fisher scientific | S369-500 | |
| Synergy HT microplate reader | BioTek | Not available | |
| Tris Base | Fisher scientific | BP152-500 |

