A Technique to Isolate Nuclei from Non-neuronal Cells in the Hippocampal Dentate Gyrus
Abstract
Source: Kerloch, T., et al. Fluorescence-Activated Nuclei Negative Sorting of Neurons Combined with Single Nuclei RNA Sequencing to Study the Hippocampal Neurogenic Niche. J. Vis. Exp. (2022).
The video demonstrates a method for isolating intact nuclei from glial and stem cells in the mouse hippocampal dentate gyrus (DG). The DG tissue is mechanically dissociated in a cold homogenization buffer to release the nuclei. Tissue debris and other released organelles are discarded, and the nuclei are stained with a DNA stain and a fluorophore-conjugated antibody specific for the neuronal nuclear antigen (NeuN). Fluorescence-activated nuclei sorting is then employed to isolate the NeuN-negative nuclei of non-neuronal cells.
Protocol
All procedures involving sample collection have been performed in accordance with the institute's IRB guidelines
1. Dissection of the DG (Timing: 15 min)
- Prepare nuclei isolation media 1 and 2 (NIM1 and NIM2), homogenization buffer (HB), and wash media (WM) (Table 1). Place all the buffers, media, reagents, and tools on ice until needed. Put the Dounce homogenizer (see Table of Materials) on ice during preparation (minimum 1 h before the homogenization step).
NOTE: NIM1 can be prepared and stored at 4 °C for up to 6 months. NIM2, HB, and WM should be freshly prepared.
CAUTION: Manipulate dithiothreitol (DTT), the protease inhibitor and Triton X-100 with care. These compounds are skin and eye irritating, acutely toxic, and hazardous for the aquatic environment. While using these chemicals wear protective gloves, clothing, eye and face protection, wash hands thoroughly after handling, and avoid release to the environment. - Euthanize a 22-month-old male C57Bl/6J mouse by cervical dislocation following the Home Office Schedule 1 procedure.
NOTE: See discussion for rationale regarding the use of 22-month-old mouse in this study. However, this protocol can be performed at any age across the lifespan. - Dissect the brain out from an euthanized mouse and transfer it into a 10 cm Petri dish filled with ice-cold 1x phosphate-buffered saline (PBS) (Figure 1). Place the Petri dish on ice. Remove the cerebellum using a scalpel and cut the brain in half between both the hemispheres (along the sagittal axis).
- Fill in a new 10 cm Petri dish with ice-cold PBS and place it on ice. Transfer one half of the brain to the new Petri dish. Using binoculars, dissect out the DG and repeat this step to obtain the second DG from the second half of the brain.
NOTE: These steps (steps 1.2-1.4) were adapted from a previously described procedure. It is important to proceed as rapidly as possible at this stage to preserve the integrity of the cells. - Transfer the two DGs into the precooled Dounce homogenizer and add 1 mL of cold HB.
2. Tissue dissociation, single nuclei isolation, and anti-NeuN immunostaining (Timing: 2 h)
- Homogenize the tissue with 10 strokes of the loose "A" pestle, followed by 15 strokes of the tight "B" pestle.
NOTE: Dounce homogenization should be performed with the mortar on ice with gentle strokes to reduce heat caused by friction and foaming. All the buffers and equipment should be precooled and kept on ice during the procedure. - Transfer the homogenate into a prechilled 15 mL tube; rinse the Dounce homogenizer with 1 mL of cold HB and combine to the same tube. Add 3 mL of HB to the 15 mL tube and incubate 5 min on ice. Mix 2x by inverting the tube gently.
- Pre-wet a 70 µm strainer cap with 0.5 mL of HB over a 50 mL test tube. Strain the nuclei suspension from step 2.2 by tipping over the 15 mL tube gently into the cell strainer. Wash the cell strainer with 0.5 mL of HB.
- Remove the cell strainer and centrifuge the test tube at 500 x g for 5 min at 4 °C, using a swing bucket centrifuge. Discard the supernatant.
NOTE: Straining the homogenate will help reduce the debris, which is critical for the flow cytometry and the single nuclei RNA sequencing (snRNA-seq) downstream steps. - Resuspend the pellet gently in 4 mL of HB using a P1000 pipette. Incubate on ice for 5 min. Spin at 500 x g for 10 min at 4 °C. Discard the supernatant and resuspend the pellet in 3 mL of WM.
- Pre-wet a 35 µm strainer cap over a 15 mL test tube with 0.5 mL of WM. Strain the nuclei suspension from step 2.5 through the cell strainer, gently pipetting 0.5 mL at a time using a P1000 pipette.
- Wash the strainer cap with 0.5 mL of WM and place the tube on ice. Transfer the filtrate to a new 15 mL tube and centrifuge for 5 min and 4 °C at 500 x g. Discard the supernatant and resuspend the pellet in 3 mL of WM.
- Spin at 500 x g for 5 min at 4 °C. Discard the supernatant and resuspend the pellet in 1 mL of WM with the mouse anti-neuronal nuclear antigen (NeuN), AlexaFluor 488-conjugated antibody (anti-NeuN-AF488, 1:32,000) and 1 µg/mL 4′,6-diamidino-2-phenylindole (DAPI). Incubate for 45 min on ice in the dark.
NOTE: To optimize the immunostaining of isolated nuclei, it is recommended to titrate the antibody to determine the optimal dilution for flow cytometry analysis and sorting. Then, run adequate controls to confirm that the staining conditions are optimal. For instance, with the conjugated anti-NeuN-AF488 antibody, a negative control (i.e., no addition of the antibody, Figure 2A) and a positive control (i.e., staining with the antibody, Figure 2B) were run to assess segregation of unstained and stained populations. When starting to work with an AF488-conjugated antibody, running an AF488-conjugated isotype control is recommended to assess specificity. If a non-conjugated antibody is used, additional control such as adding a secondary antibody only to a nuclei preparation might be necessary to assess unspecific binding of the secondary antibody.
3. Fluorescence-activated nuclei sorting (FANS) to exclude neuronal populations (Timing: 45 min)
- Transfer the immuno-stained nuclei suspension to a 5 mL test tube and keep it on ice until the start of the flow cytometry procedure.
NOTE: If working with bigger pieces of tissue than two mouse DG, further dilution with WM buffer might be required to avoid clogging the fluorescence-activated cell sorter (FACS) if nuclei density in the solution becomes high. - Vortex the samples for 3 s at low speed before placing the tubes into the FACS instrument (see Table of Materials).
NOTE: (FACS setup) Sorting machines need to be aligned at the start of the procedure with calibration particles following the manufacturer's recommendations. The drop delay was calibrated with beads or microspheres (see Table of Materials) according to the FACS model. Samples were sorted at 4 °C on purity mode. To reduce the collection volume, the nuclei were sorted through a 70 µm nozzle at the recommended pressure for the flow cytometer. Nuclei were sorted into 1.5 mL low binding tubes (see Table of Materials) containing 50 µL of WM. All collection tubes were coated in PBS + 5% bovine serum albumin (BSA) at 4 °C overnight to reduce the risk of nuclei sticking to the walls of the tube. - To acquire the data from a sample of the stained nuclei suspension, set the gates in DAPI-height and the DAPI-area in order to exclude the cell debris and the aggregated nuclei (Figure 3A). Furthermore, separate single nuclei from any remaining DAPI-stained aggregates or cell debris by setting the gates in the log side scatter (SSC)-area and the log forward scatter (FSC)-area (Figure 3B).
- Set the gates for the anti-NeuN-AF488 and the FSC-area, to isolate the NeuN-AF488-negative population, as shown in Figure 3C.
- After analysis, using the gating strategy described above, sort the NeuN-AF488-negative population in a 1.5 mL collection tube filled with 50 µL of WM.
NOTE: Following the gating strategy described above and the dissection procedure to isolate the DG from the brain of an adult mouse, the NeuN-AF488-negative population is expected to represent ~14% of the single nuclei.
Table 1: Compositions of media and buffers used in the study.
| Nuclei isolation medium #1 (NIM1) – store up to 6 months at 4 °C | ||
| Component | Volume (µL) | Final concentration (mM) |
| 1.5 M Sucrose | 1,833.30 | 250 |
| 1 M KCl | 275 | 25 |
| 1 M MgCl2 | 55 | 5 |
| 1 M Tris buffer, pH 8.0 | 110 | 10 |
| Nuclease-free water | 8,726.70 | – |
| Total volume | 11,000 | – |
| Nuclei isolation medium #2 (NIM2) – make fresh | ||
| Component | Volume (µL) | Final concentration |
| NIM1 | 10,769 | – |
| 1 mM DTT | 11 | 1 µM |
| 100x Protease inhibitor | 110 | 1x |
| Triton X-100 10% (v/v) | 110 | 0.1% (v/v) |
| Total volume | 11,000 | – |
| Homogenization buffer (HB) – make fresh | ||
| Component | Volume (µL) | Final concentration |
| NIM2 | 10,958.75 | – |
| RNase Inhibitor, 40 U µL-1 | 27.5 | 100 U mL-1 |
| RNasin, 40 U µL-1 | 13.75 | 50 U mL-1 |
| Total volume | 11,000 | – |
| Wash Media (WM) | ||
| Component | Volume (µL) | Final concentration |
| BSA 7.5% | 1,466.70 | 1% |
| RNase Inhibitor, 40 U µL-1 | 27.5 | 100 U mL-1 |
| RNasin, 40 U µL-1 | 13.75 | 50 U mL-1 |
| PBS | 9,492.05 | – |
| Total volume | 11,000 | – |
Representative Results
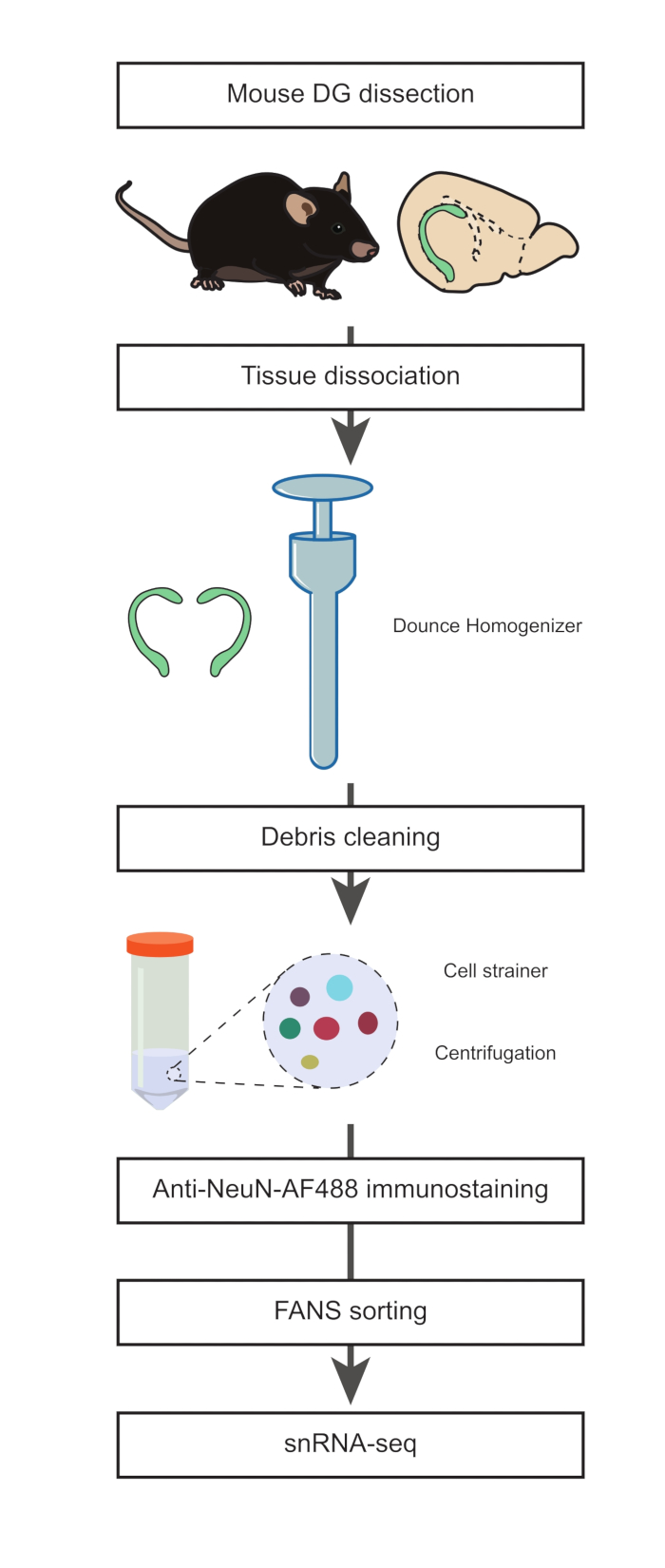
Figure 1: Preparation of a single nuclei suspension from the dissected DG of adult mice for snRNA-seq of non-neuronal populations. Flow diagram describing the main steps of the protocol that include dissection of mouse DG, preparation of a suspension of single nuclei, NeuN immunostaining, and negative NeuN-FANS-sorting before proceeding with snRNA-seq.
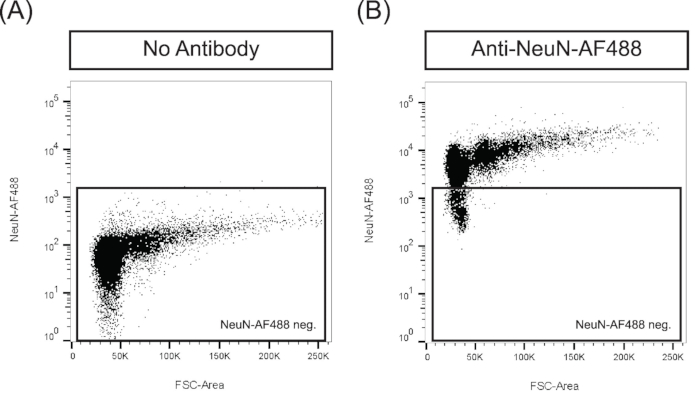
Figure 2: Validation of the immunostaining for FANS. Nuclei suspension was incubated (A) without the anti-NeuN-AF 488 antibody as a negative control or (B) with the antibody and run through the FACS sorter to validate immunostaining conditions.
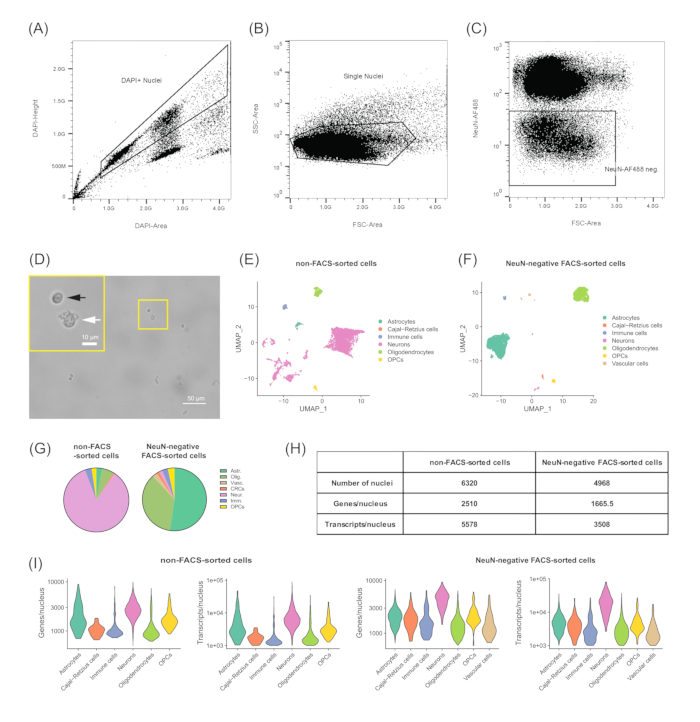
Figure 3: Gene expression analysis and re-clustering of the astrocyte cluster. (A) Uniform Manifold Approximation and Projection for Dimension Reduction (UMAP) plot showing the clustering of 4968 nuclei based on genome-wide expression profiles. Cell-type calls were done based on cell-type markers. (B) Astrocyte cluster comprised of 2579 nuclei chosen from (A) for further sub-setting to investigate potential cellular sub-types. Four sub-types were detected by Seurat (0-3) clustering, shown by different colors. (C) Gene expression levels of specific cellular markers across the four cell-types. All plots were obtained using the Seurat R package. Briefly, RNA-seq counts were normalized for each cell by the total expression and multiplied by the scale factor (10,000). This result was then log transformed. The transformed values were scaled (variance scaled to one) and centered (mean set to zero) within each cell before Uniform Manifold Approximation and Projection for Dimension Reduction (UMAP) was applied to calculate the embeddings, which were used as values on x and y axes. Graphs represent the output of a dimensional reduction technique on a 2D scatter plot where each point represents a cell with respective x and y co-ordinates based on the cell embeddings determined by the reduction technique. Cells with similar gene signatures are positioned close to one another by the embeddings.
Divulgations
The authors have nothing to disclose.
Materials
| 0.5ml microtube | Eppendorf | 30124537 | |
| 10.00µm Flouresbrite YG Carboxylate Microspheres | Polysciences | 15700-10 | |
| 15 mL polypropylene centrifuge tubes | Corning | 430052 | |
| 2 pairs of sterile Dumont #5 forceps | Fine Science Tools | 11252-30 | |
| 4′,6-diamidino-2-phenylindole (DAPI) | Sigma Aldrich | D9564-10MG | |
| 4150 TapeStation System | Agilent | N/A | |
| 5 mL round bottom high clarity polypropylene test tube with snap cap | Falcon | 352063 | |
| 5 mL round bottom polystyrene test tube with cell strainer snap cap | Falcon | 352235 | |
| 50 mL polypropylene centrifuge tubes | Corning | 430829 | |
| 70 µm cell strainer | Falcon | 352350 | |
| 8 peak SPHERO Rainbow Calibration Particles | BD Biosciences | RCP-30-5A | |
| Accudrop Beads | BD Biosciences | N/A | |
| Allegra X-30R Centrifuge | Beckman Coulter | N/A | |
| Anti-NeuN antibody, clone A60, Alexa Fluor 488 conjugated | Millipore | MAB377X | |
| BD FACSAria Fusion Flow Cytometer | BD Biosciences | N/A | |
| Beckman Coulter MoFlo XDP | Beckman Coulter | N/A | |
| BSA 7.5% | Gibco | 15260037 | |
| Dithiothreitol (DTT) | Thermo Scientific | R0861 | |
| Dounce tissue grinder set: mortar, loose pestle (A) and tight pestle (B) | KIMBLE | D8938-1SET | |
| Eppendorf Tubes Protein LoBind 1.5ml | Eppendorf | 30108116 | |
| Halt, 100x Protease inhibitor | ThermoFisher | 78429 | |
| KCl | Any chemical supplier | Laboratory made | |
| LUNA-FX7 Automated Cell counter | Logos Biosystems | N/A | |
| MgCl2 | Any chemical supplier | Laboratory made | |
| N°10 guarded sterile disposable scalpels | Swann-Morton | 6601 | |
| Nuclease-free water | Sigma Aldrich | W4502-1L | |
| Pair of sterile student surgical scissors | Fine Science Tools | 91401-12 | |
| PBS | Any chemical supplier | Laboratory made | |
| RNase Inhibitor 40 U µl-1 | Ambion | AM2684 | |
| RNasin 40 U µl-1 | Promega | N211A | |
| Sterile Petri dish | Corning | 430167 | |
| Sucrose | Sigma Aldrich | 59378-500G | |
| Tris buffer, pH 8.0 | Any chemical supplier | Laboratory made | |
| Triton X-100 10% (v/v) | Sigma Aldrich | T8787-250ML | |
| Trypan blue | Invitrogen | T10282 |

