Live-Cell Fluorescence Microscopy to Investigate Subcellular Protein Localization and Cell Morphology Changes in Bacteria
Özet
This article provides a step-by-step guide to investigate protein subcellular localization dynamics and to monitor morphological changes using high-resolution fluorescence microscopy in Bacillus subtilis and Staphylococcus aureus.
Abstract
Investigations of factors influencing cell division and cell shape in bacteria are commonly performed in conjunction with high-resolution fluorescence microscopy as observations made at a population level may not truly reflect what occurs at a single cell level. Live-cell timelapse microscopy allows investigators to monitor the changes in cell division or cell morphology which provide valuable insights regarding subcellular localization of proteins and timing of gene expression, as it happens, to potentially aid in answering important biological questions. Here, we describe our protocol to monitor phenotypic changes in Bacillus subtilis and Staphylococcus aureus using a high-resolution deconvolution microscope. The objective of this report is to provide a simple and clear protocol that can be adopted by other investigators interested in conducting fluorescence microscopy experiments to study different biological processes in bacteria as well as other organisms.
Introduction
The field of bacterial cell biology has been significantly enhanced by recent advancements in microscopy techniques1,2. Among other instruments, microscopes that are capable of conducting timelapse fluorescence microscopy experiments remain a valuable tool. Investigators can monitor various physiological events in real-time using fluorescent proteins such as, green fluorescent protein (GFP)-based transcriptional and translational reporter fusions, fluorescent D-amino acids (FDAA)3, or use other stains for labeling the cell wall, membrane and DNA. It is therefore of no surprise that fluorescence microscopy remains popular among microbial cell biologists. In addition to simply showing the end phenotypes, providing information as to how the observed phenotypes arise using timelapse microscopy could add significant value to the findings and potentially offer clues as to what cellular processes are being targeted by potential drug candidates4.
The protocols to conduct high-resolution imaging using a fully motorized, inverted, wide-field fluorescence microscope (see the Table of Materials) are provided in this article. These protocols could be adapted to suit the needs of other fluorescence microscopes that are capable of conducting timelapse microscopy. Although the software discussed here corresponds to the specific manufacturer-supplied software as indicated in the Table of Materials, software commonly supplied by other microscope manufacturers or the freely available ImageJ5, have equivalent tools for analyzing microscopy data. For conditions where timelapse is not conducive, time-course experiments could be conducted as described in this article. The protocols described here provide a detailed guide to study the phenotypic changes in two different bacterial species: B. subtilis and S. aureus. See Table 1 for strains used.
Protocol
1. General growth conditions
- Inoculate 2 mL of appropriate growth medium supplemented with antibiotics (where required) with a single colony of the strain(s) to be imaged. Incubate these seed cultures overnight at 22 °C in a shaking incubator.
NOTE: The specific bacterial growth conditions used in this article are provided under the representative results section. - Dilute overnight cultures 1:20 in fresh media in a 125 mL flask, supplement with antibiotics (where required).
- Grow culture(s) at 37 °C in a shaking incubator until mid-logarithmic phase (OD600 = 0.5).
NOTE: Inducer or inhibitor could be added directly to growing culture(s) at the appropriate growth phase. - Harvest cells at desired culture conditions for microscopy sample preparation as described in the following section.
2. Sample preparation
- Prepare 1% agarose by combining 0.25 g of molecular biology grade, low EEO, agarose with either 25 mL of growth medium supplemented with appropriate antibiotics (where required) for timelapse microscopy or sterile water. Heat the mixture with the help of a microwave for approximately 30 s and pour it into a sterile 100 mm x 15 mm Petri dish and let it solidify.
- Take a 5–50 µL aliquot of the cell culture depending on the need and stain the cells. Stain cells with appropriate dyes at this stage (e.g., add fluorescent dye FM4-64 (membrane stain) for an experiment (Figure 1)).
- Pipette a 5 µL aliquot of the culture sample or stained sample to be imaged onto the bottom of a 35 mm glass bottom culture dish (with 14 mm microwell diameter and uncoated No. 1.5 coverslip; see the Table of Materials).
CAUTION: Do not use poly-L-lysine coated coverslips especially for timelapse microscopy as they could induce different growth pattern (we have observed this with poly-D-lysine as well; data not shown) and/or affect protein dynamics6. - To minimize the use of dyes, stain 3–4 µL cell suspension (harvested at OD600 = 0.5; approximately 2.7 x 107 and 3.4 x 107 for B. subtilis and S. aureus respectively per 1 mL culture) directly on the glass bottom dish.
- Place a pre-cut agarose slab (11 mm in diameter or of any desired size to overlap the area of the coverslip; cut using the open end of a sterile tube or a razor blade) on top of the sample and gently tap to make sure the agarose slab is lying flat against the coverslip.
- Pipette a 5 µL aliquot of the culture sample or stained sample to be imaged onto the bottom of a 35 mm glass bottom culture dish (with 14 mm microwell diameter and uncoated No. 1.5 coverslip; see the Table of Materials).
- Subsequent to imaging (see following section), if the number of cells in the field of view are not desirable, adjust the cell density of the sample via dilution or concentration by centrifugation and alter the ratio of cells in the sample using growth medium/buffer as necessary.
NOTE: To minimize autofluorescence emanating from the culture medium, cells can be washed in buffer (for example in standard 1x phosphate buffer saline) and resuspended in appropriate amount of buffer prior to imaging. It is possible that in this step smaller cells or vesicles may not be retained after centrifugation and will hence be lost. - Add water using a pipette (~5 µL drops) to the space inside the culture dish (around the coverslip) to prevent the agarose pad from drying, and to help maintain humidity during the course of image acquisition, especially for timelapse experiments.
- Allow culture dishes to equilibrate to the temperature inside the incubation chamber, a built-in opaque compartment provided by the manufacturer, of the microscope for 15–20 min.
NOTE: Turn on the heating element and set the incubation chamber to a desired temperature several hours prior to imaging. This will ensure that the hardware stabilizes to the new temperature. For prolonged timelapse imaging, place a beaker or flask with water inside the microscope chamber, away from the working area, to maintain humidity.
3. Imaging
- On the day of the experiment, turn on the microscope system. Start the imaging software (see Table of Materials) by clicking the appropriate icon on the desktop. Initialize the microscope by clicking the Initialize Microscope option (button depicted with a microscope on it) on the software’s start-up dialog box. Ensure that the objective is fully lowered using the microscope coarse adjustment prior to initialization.
NOTE: Following initialization three additional dialog boxes should appear in addition to the start menu: resolve3D, data collection, and filter monitor. - Place a drop of 1.517 (refractive index) oil on the 100x oil immersion objective supplied by the manufacturer (Numerical Aperture = 1.4, Working Distance = 0.12 mm; see Table of Materials).
NOTE: It is important to choose the appropriate immersion oil for the temperature at which imaging is conducted. - Load the glass bottom dish containing sample into the metal housing (coffin) and gently slide into the stage clamp.
- Use the coarse adjustment knob to raise the objective until the oil makes contact with the glass bottom of the dish. Use the eye piece and fine adjustment knob to bring the sample into focus. Once the cells are in focus turn the knob from the eye piece to camera mode by moving the selector switch located on the front of the microscope body to the left.
- Begin experiment using the imaging software. On the resolve3D window, select the Design/run experiment icon depicted by a flask. A new dialog box should appear entitled design/run experiment.
- Set the number of Z-stacks and sample thickness using the design and then sectioning tab on the design/run experiment dialog box.
NOTE: For the experiments in representative results section, 17 Z-stacks at 200 nm interval for still images and four Z-stacks at 200 nm interval for timelapse microscopy were used. - To measure the thickness of the cells in the sample, manually adjust the Z-plane incrementally by using the up and down arrows on the resolve3D dialog box. Mark where the cells go out-of-focus as the upper and lower limit for image acquisition. Import this information prior to running the experiment.
- To help minimize phototoxicity and photobleaching during timelapse imaging, reduce the number of Z-stacks and choose the mid-plane of the cells for image acquisition.
- Set the number of Z-stacks and sample thickness using the design and then sectioning tab on the design/run experiment dialog box.
- Select the appropriate filter set for the experiment using the design and then channels tab on the design/run experiment dialog box (TRITC: EX 542/27; EM 597/45; FITC/GFP: EX 475/28; EM 525/48; mCherry: EX 575/25; EM 632/60; Cy5: EX 632/22; EM 676/34).
- Also, select a reference for the collection of POL/DIC information. Adjust percentage transmission (light intensity) and duration of exposure for individual channels selected prior to imaging by selecting the appropriate options on the resolve3D dialog box.
NOTE: In a test field of view that is not considered for the experiment test these settings to identify if the selected parameters obtain meaningful fluorescence data without missing weak signal or oversaturating it. Then import these parameters for the experimental set up.
- Also, select a reference for the collection of POL/DIC information. Adjust percentage transmission (light intensity) and duration of exposure for individual channels selected prior to imaging by selecting the appropriate options on the resolve3D dialog box.
- Open the points list by selecting the Points list button on the resolve3D dialog box. A new dialog box should appear entitled points list. Mark several fields of view to be used in the experiment by finding appropriate fields of view using the microscope stage controls and selecting the mark point option on the points list dialog box.
NOTE: A replace point will have to be selected each time the microscope is refocused on a point in the points list. This can be done by focusing the microscope on the appropriate point in the list and then selecting the replace point button. It is important to not touch the analog coarse/fine adjustment knob when refocusing; use only the software to adjust the focus. - Set timelapse parameters by first selecting the design and then timelapse tab on the design/run experiment dialog box. Select the timelapse check box. Enter the appropriate timelapse parameters in terms of timelapse images/total time.
- Set points to be imaged from the points list by first selecting the design and then points tab on the design/run experiment dialog box. Select the visit points list option and enter points to be imaged in the text box separated by commas or hyphens if it is a complete sequence.
- Prior to beginning an experiment, edit file names and file locations using the run tab on the design/run dialog box. File location can be changed by selecting the settings button, selecting the data folder, and by then selecting the appropriate folder or creating a new one. Change file names by entering in the new file name in the image file name text box.
- Begin the experiment by selecting the play button on the begin experiment dialog box.
NOTE: For timelapse, continually check focus at each field of view throughout the experiment using DIC setting (to avoid unnecessary photobleaching) and refocus and update the information in points list as cells tend to go out of focus over time.
4. Image processing
- Open the desired raw (R3D) image files for generation of figures.
NOTE: Recently saved files can be located using the data folder button on the imaging software’s start-up dialog box. - Run the deconvolution program, to remove out-of-focus fluorescence light7,8, in order to produce a deconvolved D3D image file. Select the process tab on the start-up dialog box, and then select deconvolve. A new dialog box will appear entitled deconvolve.
- Set the deconvolution parameters by dragging the appropriate image number (located in the upper left-hand corner of each image file) to the input on the deconvolution dialog box. On the deconvolution dialog box, select the more options button, and deselect the crop border rolloff after processing check box (otherwise the images cannot be superimposed over the corresponding DIC file).
- Click the do it button on the deconvolution dialog box to deconvolve raw image file.
NOTE: The deconvolution parameters could be adjusted as indicated by the software manufacturer’s guidance manual for desired results.
- If necessary, perform manual background noise subtraction and brightness/contrast adjustment in any or all wavelength (color) channels. Do this by selecting the contrast adjustment button on the deconvolved image file.
- Save the image as a TIFF file by first selecting the file option at the top of the D3D dialog box. Then select save as TIFF, identify and select appropriate Z-stacks (that are within focus), select or unselect desired filter sets.
- Perform data quantification using any manufacturer-supplied software or freely available programs such as ImageJ5.
- For cell size quantification click on the tool tab on the appropriate D3D image file, and then select the measure distances option. Measure distances by selecting start and end points on the desired image file using the mouse through left clicks.
NOTE: It is best to zoom in on the image during cell size quantification. - For fluorescence signal quantification use the data inspector option under the tool tab on the appropriate D3D image file.
NOTE: It is best to zoom in on the image and to have only relevant filters selected when completing fluorescence signal quantification. - To quantify the fluorescence signal, draw a box with column/row option set to specific dimension. Select an area with signal to be quantified and record the value. Measure the background signal by using the same size box to select an area immediately outside of the cells. Subtract the background value from the value recorded from the fluorescence signal obtained within the cell.
- For cell size quantification click on the tool tab on the appropriate D3D image file, and then select the measure distances option. Measure distances by selecting start and end points on the desired image file using the mouse through left clicks.
Representative Results
GpsB phenotypes
Previously we have shown that Sa-GpsB is an essential protein as depletion of GpsB using an antisense RNA results in cell lysis9. Here we describe how the emergence of various cell division phenotypes and changes in protein localization could be captured using the timelapse microscopy protocol described in this article. For this purpose, S. aureus strains RB143 [SH1000 harboring pEPSA5 (empty vector)] and GGS8 [SH1000 harboring pGG59 (Pxyl-gpsBantisense bla cat)] reported previously9, were grown as follows. Strains RB143 and GGS8 were inoculated in 2 mL of tryptic soy broth (TSB) supplemented with 5 µg/mL chloramphenicol (chlor) in a 15 mL test tube and were incubated overnight at 22 °C while shaking. The overnight cultures were diluted 1:20 in 10 mL of fresh TSB + chlor in a 125 mL flask and grown at 37 °C with shaking until mid-logarithmic phase (OD600 = 0.5). The inducer, 1% xylose, was added to the culture medium to trigger the expression of antisense RNA of gpsB and the culture was grown for another 3 h. Cells were then stained with fluorescent dye FM4-64 (membrane stain), where required, by the addition of 0.5 µL of a 10 µg/mL stock of FM4-64 directly onto the 5 µL aliquot of culture on the microscope dish as described in the protocol section. As shown in Figure 1 and Video 1, addition of xylose to induce GGS8 strain resulted in a “sick” cell phenotype, as described previously9, while empty vector control (RB143) appeared similar to our control—cells grown in the absence of inducer.
Our group also reported that overproduction of S. aureus GpsB (Sa-GpsB) disrupts cell division in B. subtilis9. We use this overexpression phenotype as an example to demonstrate the protocol described here. To this end, a B. subtilis strain GG9 (amyE::Phyperspank-gpsBSa spc; ftsAZ::ftsAZ-gfpΩerm) was used9. Subcellular localization of fluorescently-labeled FtsZ, a key cell division protein which marks the cell division sites10,11, was used to monitor the status of cell division. The sample for microscopy was prepared as follows. A single colony of GG9 was inoculated in 2 mL Luria-Bertani (LB) medium and incubated overnight at 22 °C in an incubator shaker. The overnight cultures were 1:20 in 10 mL of fresh LB, and grown at 37 °C with shaking until mid-logarithmic phase (OD600 = 0.5). GG9 cells (5 µL aliquot) to be imaged were placed on the bottom of a glass bottom culture dish and covered with a 1% agarose pad made with LB supplemented with 250 µM (final concentration) of isopropyl β-D-1-thiogalactopyranoside (IPTG) to induce the expression of Sa-gpsB (Figure 2 and Video 2). Timelapse microscopy and cell length quantification were performed as described in the protocol section.
Inhibition of FtsZ
FtsZ, being a protein essential for cell division, is considered an attractive drug target and multiple groups are developing FtsZ inhibitors as a way to develop new antibiotics12. Localization patterns of FtsZ or one of the proteins associated with it, such as ZapA, can be used as a reporter to study and/or identify novel antimicrobial compounds. We use the protocol provided here to demonstrate this approach using S. aureus RB197 [SH1000 harboring pRB42 (PCd-zapASa-gfp bla erm)]9 and B. subtilis PE92 (ftsAZ::ftsAZ-gfpΩerm)13 strains. RB197 and PE92 strains were grown as described above in TSB (containing 5 µg/mL erythromycin; and 1.25 µM CdCl2 to induce the expression of zapA-gfp) and LB respectively. At mid-logarithmic phase, a well-characterized FtsZ inhibitor, PC19072314,15, was added at 2 µg/mL final concentration and its effect on the S. aureus and B. subtilis cells were monitored using microscopy at different time intervals (Figure 3 and Figure 4). Quantification of cell diameter of S. aureus and cell length of B. subtilis was performed as described in the protocol section.
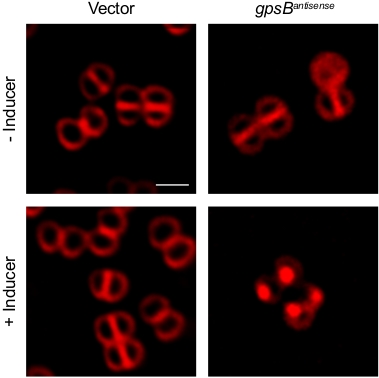
Figure 1: High-resolution micrograph of S. aureus cells displaying sick phenotype. Fluorescence micrographs of S. aureus strains harbouring either empty vector (left; RB143) or an inducible copy of antisense RNA of gpsBSa (right; GGS8) in the presence and absence of 1% xylose (inducer). Cells awere stained with FM4-64 membrane stain (stocks dissolved in sterile water) and imaged using TRITC filter set. Scale bar: 1 µm. Please click here to view a larger version of this figure.
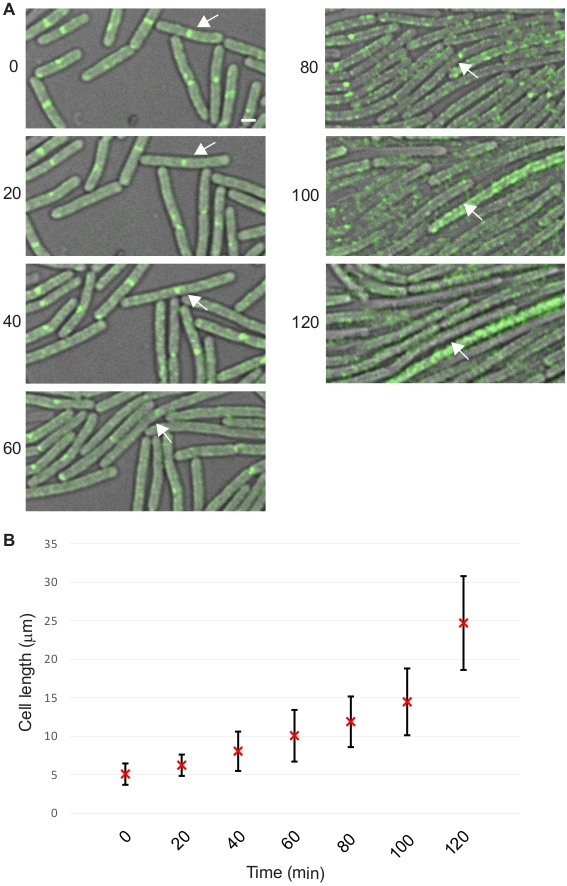
Figure 2: Representative data showing cell division inhibition in B. subtilis. (A) Timelapse micrographs of B. subtilis strain GG9 with images acquired at 20-min intervals for 120 min using the DIC/FITC channels. Fluorescence data of FtsZ-GFP (green) are shown. Arrows follow one cell throughout the experiment. Scale bar: 1 µm. (B) Quantification of cell lengths at all time points. Average cell length with error bars indicating standard deviation (n = 50) are shown. Please click here to view a larger version of this figure.
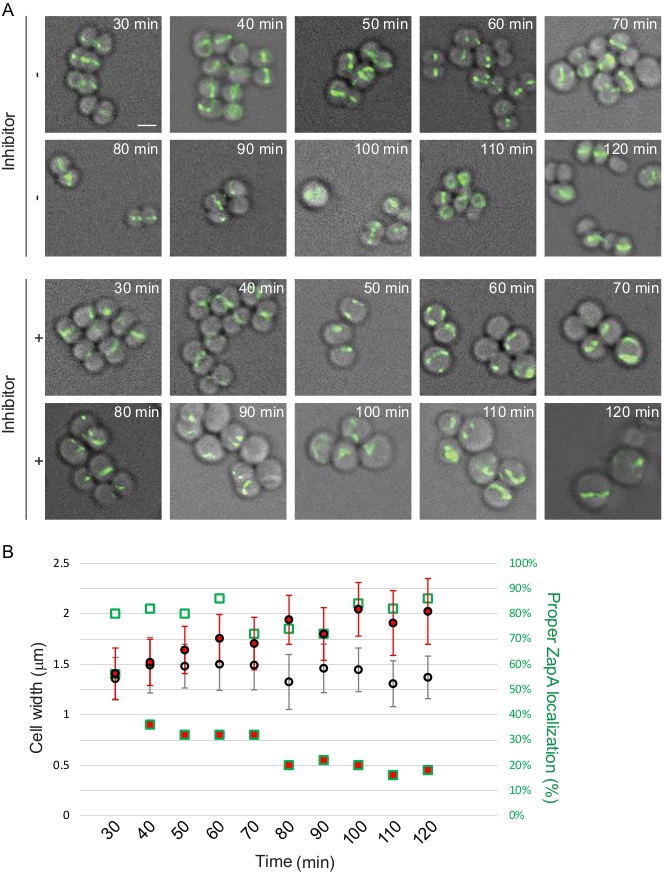
Figure 3: Time course investigation of cell division inhibition in S. aureus. (A) Mid-logarithmic phase cells of strain RB197 untreated (top) or treated (bottom) with FtsZ inhibitor (PC190723), subsequent to 30 min growth, aliquots of growing cultures were taken every 10 min for 90 min and imaged using DIC and FITC filter sets. Fluorescence from ZapA-GFP is shown. Scale bar: 1 µm. (B) Quantification of microscopy data. Average cell width with error bars indicating standard deviation (n = 50) and percentage of cells (n = 50) displaying proper ZapA-GFP localization (mid-cell and periphery) are shown. Data points of cells treated with inhibitor are shown in red. Shapes with green outline corresponds to the right Y-axis. Please click here to view a larger version of this figure.
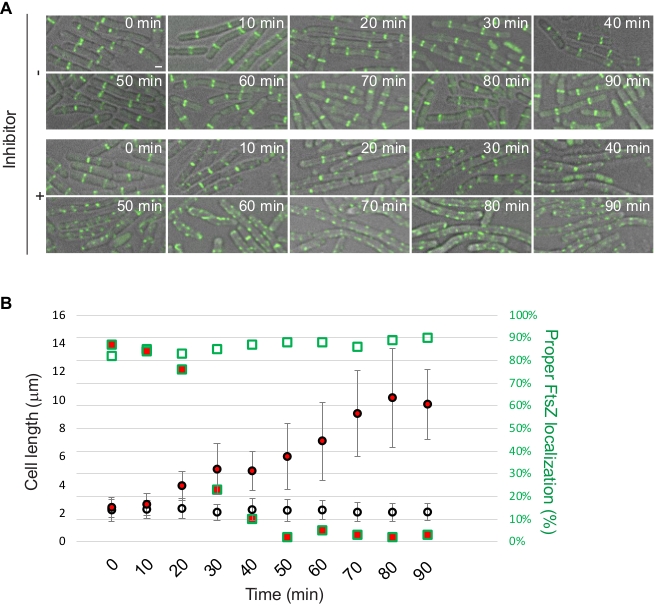
Figure 4: Investigation of cell division inhibition by a synthetic inhibitor in B. subtilis. B. subtilis strain PE92 was either untreated or treated with an FtsZ inhibitor (PC190723) at mid-logarithmic phase and were monitored for the subsequent 90 min. Aliquots of growing cultures were taken every 10 min for microscopy and images were acquired using DIC/FITC channels. Fluorescence from FtsZ-GFP is shown. Scale bar: 1 µm. (B) Quantification of microscopy data. Average cell length with error bars indicating standard deviation (n = 50) and percentage of cells (n = 50) displaying proper mid-cell FtsZ-GFP localization are shown. Data points of cells treated with inhibitor are shown in red. Shapes with green outline corresponds to the right Y-axis. Please click here to view a larger version of this figure.
Video 1: Timelapse microscopy of S. aureus cells developing sick phenotype. Strain GGS8 (gpsB antisense) treated with 1% xylose. Cells were stained with FM4-64 membrane stain and imaged at 10 min intervals for 60 min using the TRITC channel as described in the protocol. Please click here to download this video.
Video 2: Overexpression of Sa-gpsB leads to inhibition of cell division in B. subtilis. Timelapse video showing filamentation and change in FtsZ-GFP localization in GG9. Images were taken at 20 min intervals for 120 min using DIC and FITC channels. Please click here to download this video.
| Species | Strain | Genotype | Reference | |
| S. aureus | RB143 | SH1000 pEPSA5, bla, cat | Eswara et al, 2018 | |
| S. aureus | GGS8 | SH1000 pGG59 (pEPSA5 backbone) Pxyl-gpsBantisense, bla, cat | Eswara et al, 2018 | |
| S. aureus | RB197 | SH1000 pRB42 (pJB67 backbone) PCd-zapASA-gfp, bla, cat | Eswara et al, 2018 | |
| B. subtilis | GG9 | amyE::Phyperspank-gpsBSA spc; ftsAZΩftsAZ-gfp erm | Eswara et al, 2018 | |
| B. subtilis | PE92 | ftsAZ::ftsAZ-gfp Ωerm | Brzozowski et al, 2019 | |
Table 1: Strains used.
Discussion
Microscopy has remained a mainstay in studies pertaining to microbial organisms. Given their micron-scale cell size, single-cell level studies have traditionally relied on electron microscopy (EM). Although EM has become quite a powerful technique in recent years, it has its own intrinsic limitations in addition to limited user access16. Improvements in fluorescence microscopy techniques and development of different fluorescent probes, such as FDAA3, have provided microbial cell biologists with a vast array of tools to study various cellular processes in live cells. Researchers are also actively building fluorescent tools to monitor changes, for example in the level of signaling molecules such as c-di-GMP among others, in living cells17,18. In addition, high-resolution timelapse fluorescence microscopy allows investigators to monitor changes as they happen and to study relevant phenotypes.
We have provided detailed protocols to conduct microscopy experiments with a high-resolution fluorescence microscope (see Table of Materials). However, the steps in the protocols could be altered to fit the needs of the user and the microscope used. We use S. aureus and B. subtilis as our model organisms to show how to monitor various cell division phenotypes, track the changes in protein localization, and quantify the data. In addition, for cases where timelapse is not conducive, we show with the help of an FtsZ inhibitor, how to set up a time course experiment.
The inherent limitation with fluorescence microscopy is the resolution set by the diffraction limit, which could be overcome to some extent with the aid of advanced super-resolution microscopy techniques19,20. Other issues such as phototoxicity and photobleaching could be circumvented by collecting fewer Z-stacks or minimizing the duration and/or frequency of exposure to laser. Other guidance materials specific to live-cell microscopy are available21. Apart from Gram-positive organisms B. subtilis and S. aureus, using this set up, we have successfully imaged Gram-negative bacterium Escherichia coli, yeast Saccharomyces cerevisiae, and nematode Caenorhabditis elegans.
In addition to the experiments described here, similar methodologies could be used to identify compounds that target specific cellular processes in a high-throughput fashion. Algorithms that automate the quantification process can also be incorporated for large datasets22,23. There is an immense need to study different bacterial species to address the antibiotic-resistance crisis and more studies are warranted to understand the mechanisms of basic biological processes and to identify novel therapeutic compounds. Various fluorescence microscopy techniques have gained the power and momentum to aid researchers in addressing these challenges among others.
Açıklamalar
The authors have nothing to disclose.
Acknowledgements
We thank our lab members for their comments on this article. This work was funded by a start-up grant from the University of South Florida (PE).
Materials
| Agarose | Fisher BioReagents | BP160-100 | Molecular Biology Grade – Low EEO |
| DAPI | Invitrogen | D3571 | Microscopy |
| FM4-64 | Invitrogen | T3166 | Microscopy |
| Glass bottom dish | MatTek | P35G-1.5-14-C | Microscopy |
| IPTG | Fisher BioReagents | BP1755-10 | Dioxane-free |
| Microscope | GE | DeltaVision Elite | Customized Olympus IX-71 Inverted Microscope Stand; Custom Illumination Tower and Transmitted Light Illuminator Module. Objectives: PLAPON 60X (N.A. 1.42, WD 0.15 mm); OLY 100X OIL (N.A. 1.4, WD 0.12 mm); DIC Prism Nomarski for 100X Objective; CoolSnap HQ2 camera; SSI Assembly 7-color; Environmental control chamber – opaque. |
| PC190723 | MilliporeSigma | 3445805MG | FtsZ inhibitor |
| SoftWorx | GE | Manufacturer-supplied imaging software |
Referanslar
- Coltharp, C., Xiao, J. Superresolution microscopy for microbiology. Cellular Microbiology. 14 (12), 1808-1818 (2012).
- Holden, S. Probing the mechanistic principles of bacterial cell division with super-resolution microscopy. Current Opinion in Microbiology. 43, 84-91 (2018).
- Hsu, Y. P., et al. Full color palette of fluorescent d-amino acids for in situ labeling of bacterial cell walls. Chemical Science. 8 (9), 6313-6321 (2017).
- Nonejuie, P., Burkart, M., Pogliano, K., Pogliano, J. Bacterial cytological profiling rapidly identifies the cellular pathways targeted by antibacterial molecules. Proceedings of the National Academy of Sciences of the USA. 110 (40), 16169-16174 (2013).
- Rueden, C. T., et al. ImageJ2: ImageJ for the next generation of scientific image data. BioMed Central Bioinformatics. 18 (1), 529 (2017).
- Colville, K., Tompkins, N., Rutenberg, A. D., Jericho, M. H. Effects of poly(L-lysine) substrates on attached Escherichia coli bacteria. Langmuir. 26 (4), 2639-2644 (2010).
- McNally, J. G., Karpova, T., Cooper, J., Conchello, J. A. Three-dimensional imaging by deconvolution microscopy. Methods. 19 (3), 373-385 (1999).
- Wallace, W., Schaefer, L. H., Swedlow, J. R. A workingperson’s guide to deconvolution in light microscopy. Biotechniques. 31 (5), 1076-1082 (2001).
- Eswara, P. J., et al. An essential Staphylococcus aureus cell division protein directly regulates FtsZ dynamics. Elife. 7, (2018).
- Haeusser, D. P., Margolin, W. Splitsville: structural and functional insights into the dynamic bacterial Z ring. Nature Reviews Microbiology. 14 (5), 305-319 (2016).
- Du, S., Lutkenhaus, J. At the Heart of Bacterial Cytokinesis: The Z Ring. Trends in microbiology. , (2019).
- Haranahalli, K., Tong, S., Ojima, I. Recent advances in the discovery and development of antibacterial agents targeting the cell-division protein FtsZ. Bioorganic and Medicinal Chemistry. 24 (24), 6354-6369 (2016).
- Brzozowski, R. S., et al. Deciphering the Role of a SLOG Superfamily Protein YpsA in Gram-Positive Bacteria. Frontiers in Microbiology. 10, 623 (2019).
- Elsen, N. L., et al. Mechanism of action of the cell-division inhibitor PC190723: modulation of FtsZ assembly cooperativity. Journal of the American Chemical Society. 134 (30), 12342-12345 (2012).
- Haydon, D. J., et al. An inhibitor of FtsZ with potent and selective anti-staphylococcal activity. Science. 321 (5896), 1673-1675 (2008).
- Pilhofer, M., Ladinsky, M. S., McDowall, A. W., Jensen, G. J. Bacterial TEM: new insights from cryo-microscopy. Methods in Cell Biology. 96, 21-45 (2010).
- Ni, Q., Mehta, S., Zhang, J. Live-cell imaging of cell signaling using genetically encoded fluorescent reporters. Federation of European Biochemical Societies Journal. 285 (2), 203-219 (2018).
- Weiss, C. A., Hoberg, J. A., Liu, K., Tu, B. P., Winkler, W. C. Single-Cell Microscopy Reveals That Levels of Cyclic di-GMP Vary among Bacillus subtilis Subpopulations. Journal of Bacteriology. 201 (16), (2019).
- Schneider, J. P., Basler, M. Shedding light on biology of bacterial cells. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences. 371 (1707), (2016).
- Yao, Z., Carballido-Lopez, R. Fluorescence imaging for bacterial cell biology: from localization to dynamics, from ensembles to single molecules. Annual review of microbiology. 68, 459-476 (2014).
- Ettinger, A., Wittmann, T. Fluorescence live cell imaging. Methods in Cell Biology. 123, 77-94 (2014).
- Fredborg, M., et al. Automated image analysis for quantification of filamentous bacteria. BioMed Central Microbiology. 15, 255 (2015).
- Ursell, T., et al. precise quantification of bacterial cellular dimensions across a genomic-scale knockout library. BioMed Central Biology. 15 (1), 17 (2017).

